TF_ChIP-seq_ENCSR620VIC
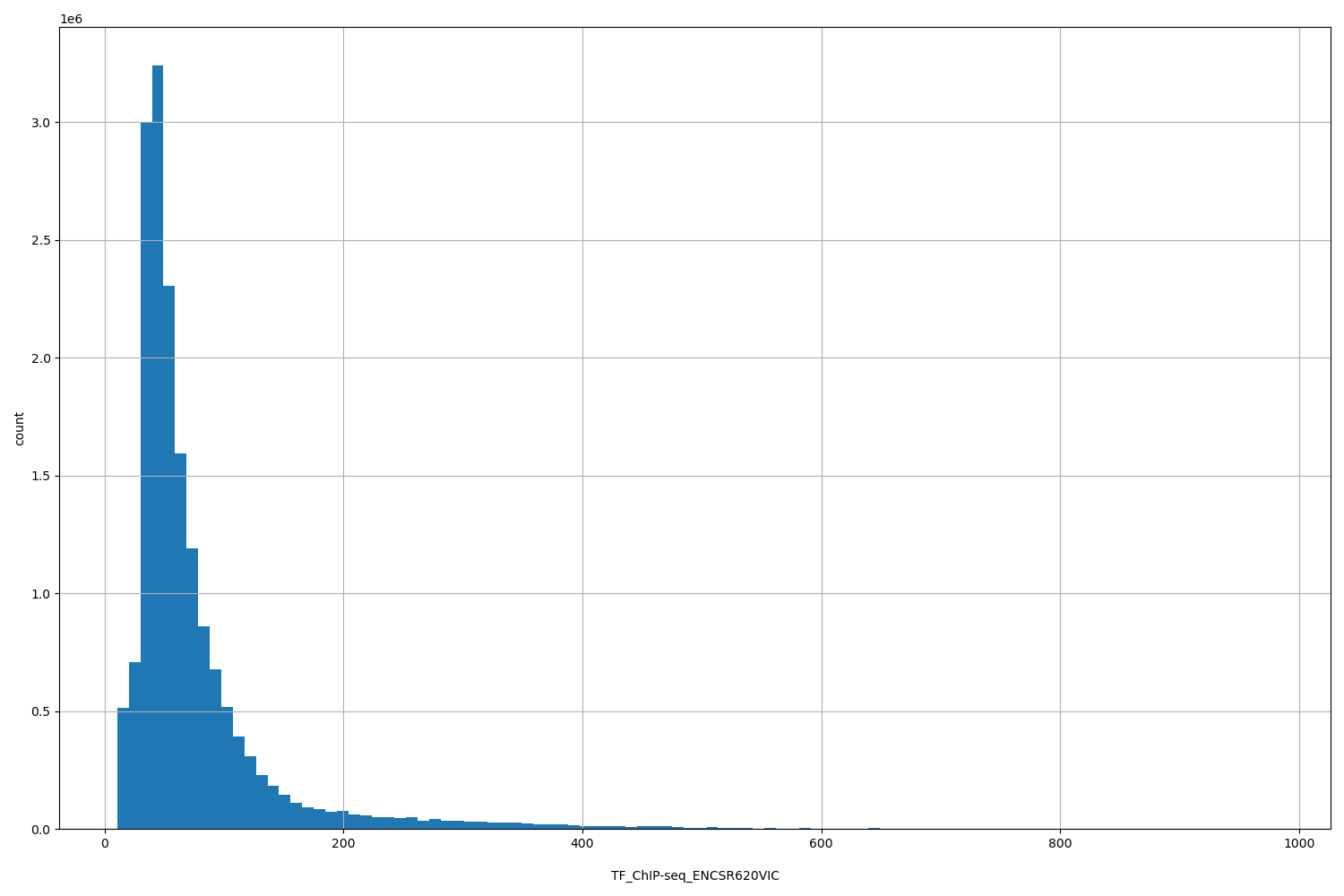
| Id: | TF_ChIP-seq/ENCSR620VIC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR620VIC [biosamplesummary="Homo sapiens K562 stably expressing CEBPG" and target="CEBPG"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: stably expressing C-terminal eGFP-tagged CEBPG output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN730FKQ|/analyses/ENCAN730FKQ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF086CSF|/files/ENCFF086CSF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.13 and a self consistency ratio of 54.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF707BHB|/files/ENCFF707BHB/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.13 and a self consistency ratio of 54.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR620VIC | float |
TF_ChIP-seq_ENCSR620VIC |
TF_ChIP-seq ENCSR620VIC [biosample_summary="Homo sapiens K562 stably expressing CEBPG" and target="CEBPG"]
|
 |
[10.6, 978] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF086CSF.bed.gz | 659.09 KB | 014da54ab1af05561f5b8bf9aa5822da |
| ENCFF086CSF.bed.gz.dvc | 100.0 B | c05c5e9aaafd0b8180ab3c9dd246ef10 |
| ENCFF086CSF.tabix.bed.gz | 500.57 KB | 12461e6a90c49591466c97644499fbf9 |
| ENCFF086CSF.tabix.bed.gz.dvc | 106.0 B | f20ffe90b489871ffb645e3b0c05541f |
| ENCFF086CSF.tabix.bed.gz.tbi | 210.69 KB | 662312ccc128f1d135f7b26466c96c2b |
| ENCFF086CSF.tabix.bed.gz.tbi.dvc | 110.0 B | 12d60e8fb1ea12282407f925971c0078 |
| genomic_resource.yaml | 2.52 KB | 7be97c48eb45d119e58f069212c3adcf |
| genomic_resource_original.yaml | 2.41 KB | 1fe09efe48185aac47cb695a68e71bb5 |
| statistics/ |