TF_ChIP-seq_ENCSR205SKQ
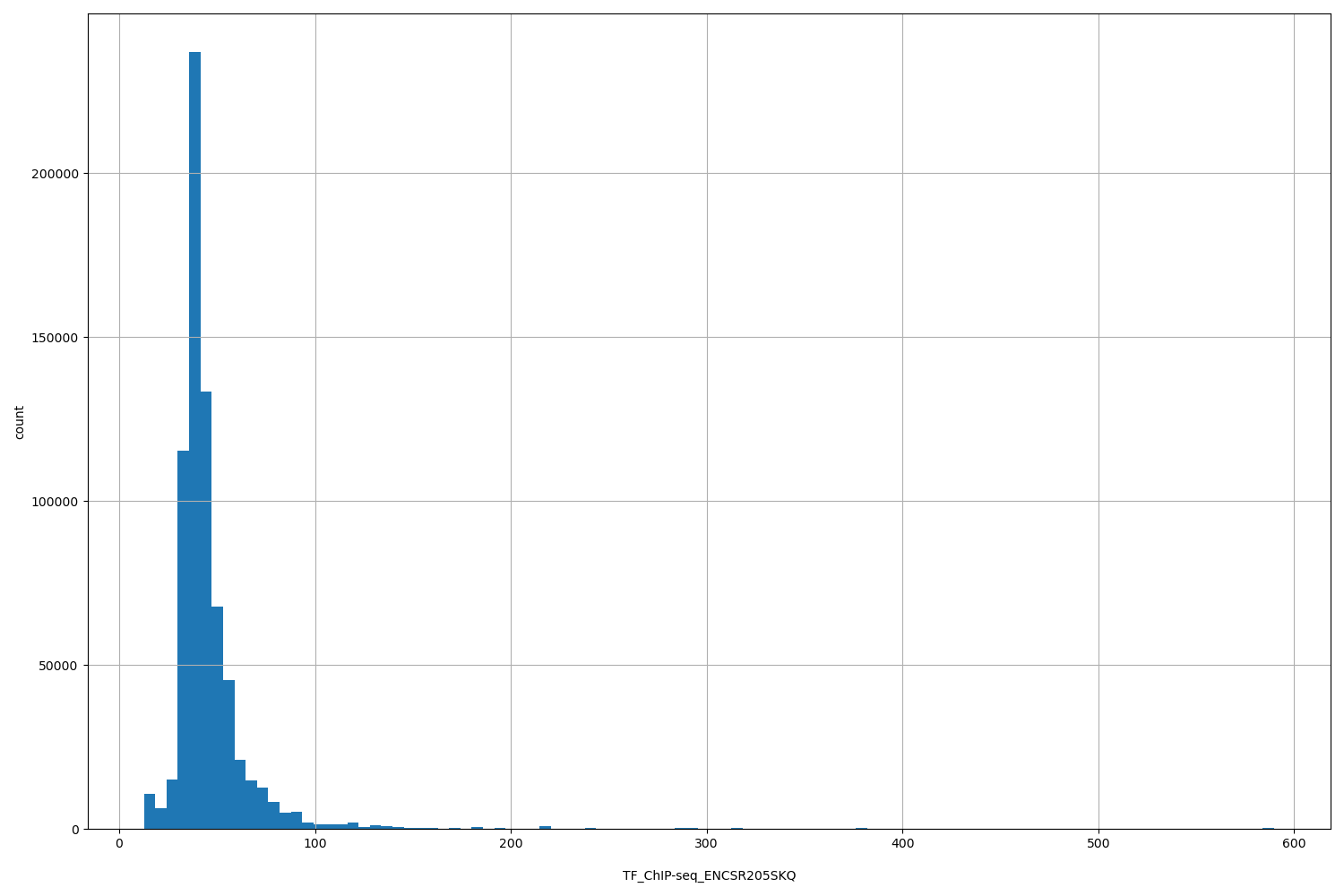
| Id: | TF_ChIP-seq/ENCSR205SKQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR205SKQ [biosamplesummary="Homo sapiens GM12878" and target="YBX1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN907ITH|/analyses/ENCAN907ITH/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF500RBO|/files/ENCFF500RBO/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Released file {ENCFF500RBO|/files/ENCFF500RBO/} has replaced subobject file {e85dd2a7-a764-4120-b41f-aec03a6234d7|/files/e85dd2a7-a764-4120-b41f-aec03a6234d7/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF699GCM|/files/ENCFF699GCM/}, {ENCFF500RBO|/files/ENCFF500RBO/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 2.47 and a self consistency ratio of 1.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF821YKR|/files/ENCFF821YKR/}, {ENCFF551ECC|/files/ENCFF551ECC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 2.47 and a self consistency ratio of 1.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR205SKQ | float |
TF_ChIP-seq_ENCSR205SKQ |
TF_ChIP-seq ENCSR205SKQ [biosample_summary="Homo sapiens GM12878" and target="YBX1"]
|
 |
[12.6, 590] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF500RBO.bed.gz | 43.15 KB | 9fe3473912e0bddc4a036473555613ba |
| ENCFF500RBO.bed.gz.dvc | 99.0 B | 93bcdbcca9a7f7486cb1799bb9c245ba |
| ENCFF500RBO.tabix.bed.gz | 27.88 KB | 13fa9b68505ba689052464a37f820b1a |
| ENCFF500RBO.tabix.bed.gz.dvc | 105.0 B | 858f7896daa0696db831b61360f45567 |
| ENCFF500RBO.tabix.bed.gz.tbi | 25.96 KB | 9337ae543333077c08982b8a0343b018 |
| ENCFF500RBO.tabix.bed.gz.tbi.dvc | 109.0 B | 52b4a8eafffe97d5c8031a90a234139e |
| genomic_resource.yaml | 2.87 KB | 7e902ea9d0b52505f81c53c1d37afe7d |
| genomic_resource_original.yaml | 2.77 KB | 73d9ff8e7fbf38b97aad48735a1966b5 |
| statistics/ |