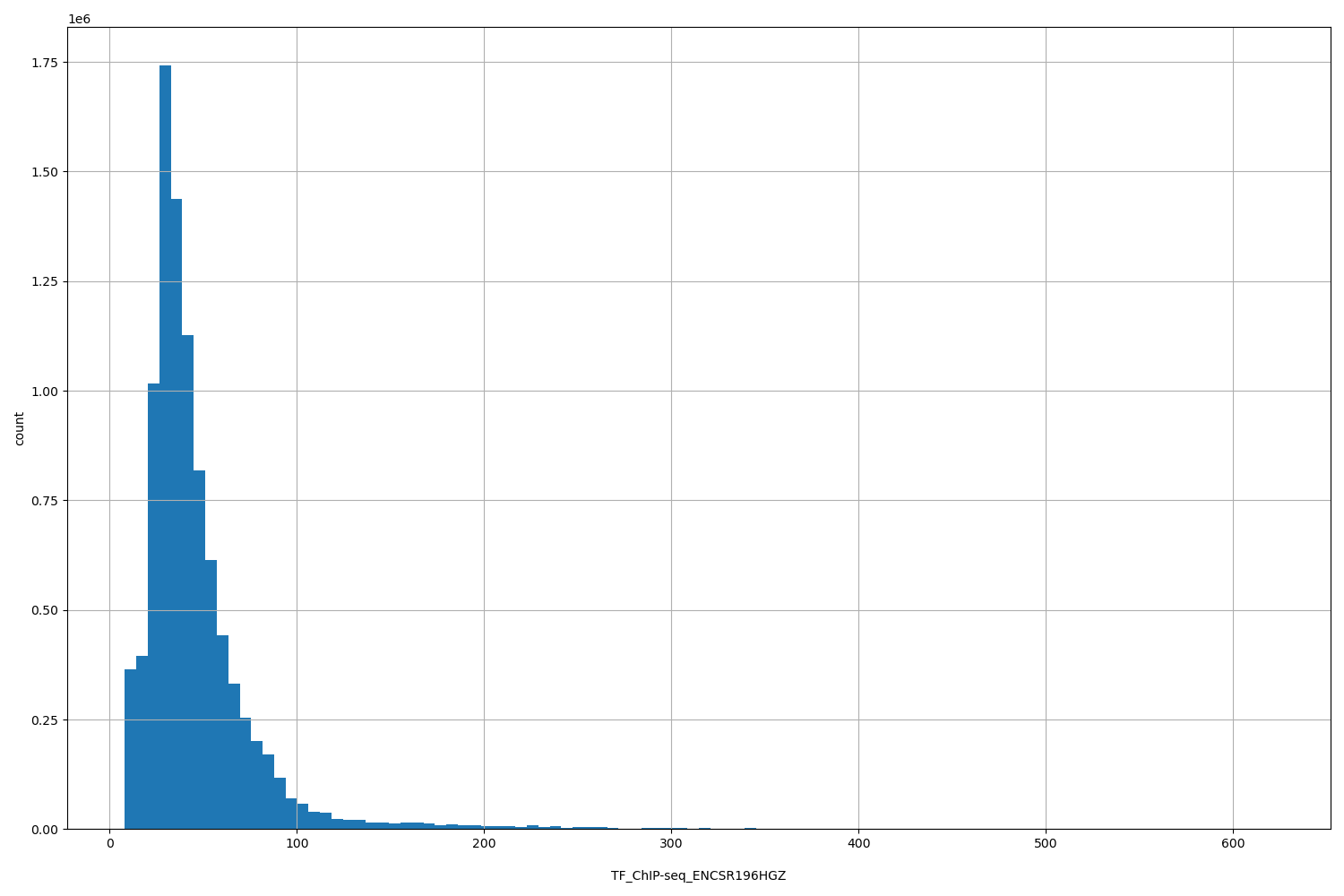
TF_ChIP-seq_ENCSR196HGZ
| Id: | TF_ChIP-seq/ENCSR196HGZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR196HGZ [biosamplesummary="Homo sapiens liver tissue female child (4 years)" and target="JUND"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female child (4 years) output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN755YYQ|/analyses/ENCAN755YYQ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF014COI|/files/ENCFF014COI/}, {ENCFF441ULY|/files/ENCFF441ULY/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.09. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF235XPK|/files/ENCFF235XPK/}, {ENCFF693FRV|/files/ENCFF693FRV/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.09. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR196HGZ | float |
TF_ChIP-seq_ENCSR196HGZ |
TF_ChIP-seq ENCSR196HGZ [biosample_summary="Homo sapiens liver tissue female child (4 years)" and target="JUND"]
|
 |
[8.16, 621] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF693FRV.bed.gz | 385.06 KB | 433d0c39ff3942ba18f059f8a4b159d1 |
| ENCFF693FRV.bed.gz.dvc | 100.0 B | ffbff85ba835098160e353bfed49c9f1 |
| ENCFF693FRV.tabix.bed.gz | 294.56 KB | f2b6f4bfc4d14fa4700c3e88260c8816 |
| ENCFF693FRV.tabix.bed.gz.dvc | 106.0 B | 3e06b3409000e97707ba42a0d476caec |
| ENCFF693FRV.tabix.bed.gz.tbi | 137.36 KB | 8df71835e3df20c6a44c6e8596d593ea |
| ENCFF693FRV.tabix.bed.gz.tbi.dvc | 110.0 B | b0083b02904c24739223414cd8a0ca3b |
| genomic_resource.yaml | 2.91 KB | 544e1b165bc1af68b9ed6abd546c7b45 |
| genomic_resource_original.yaml | 2.79 KB | 5f9a5d34e13573ee5c93535f9785b322 |
| statistics/ |