TF_ChIP-seq_ENCSR174GOO
| Id: | TF_ChIP-seq/ENCSR174GOO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR174GOO [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HMGXB4" and target="HMGXB4"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens HMGXB4 output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF883FJF|/files/ENCFF883FJF/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN053LNH|/analyses/ENCAN053LNH/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF883FJF|/files/ENCFF883FJF/}, {ENCFF429KHK|/files/ENCFF429KHK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.48 and a self consistency ratio of 2.65. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF128UTW|/files/ENCFF128UTW/}, {ENCFF250RLQ|/files/ENCFF250RLQ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.48 and a self consistency ratio of 2.65. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
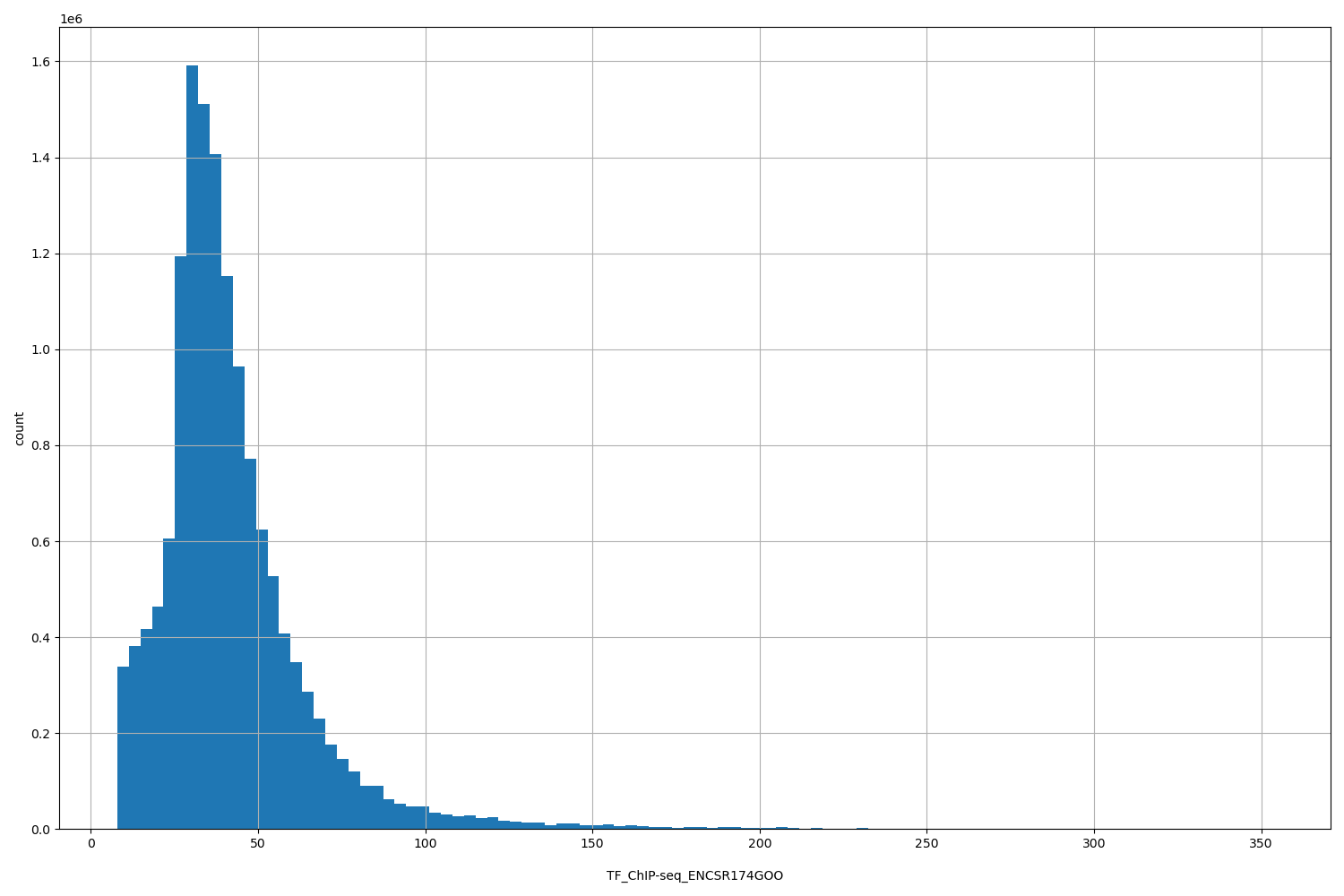
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR174GOO | float |
TF_ChIP-seq_ENCSR174GOO |
TF_ChIP-seq ENCSR174GOO [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HMGXB4" and target="HMGXB4"]
|
 |
[7.9, 353] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF883FJF.bed.gz | 539.57 KB | 1b714a75ae834b3f13b337de5efb087d |
| ENCFF883FJF.bed.gz.dvc | 100.0 B | 045203c7a745924baf7f1ab0550c0bdf |
| ENCFF883FJF.tabix.bed.gz | 425.8 KB | 8259f74f5dcf59f24ede396ff4f3751e |
| ENCFF883FJF.tabix.bed.gz.dvc | 106.0 B | a54699eb3261cfacead4e0dc8e35488d |
| ENCFF883FJF.tabix.bed.gz.tbi | 154.65 KB | 94d0c3b681ffd10298981a82172da1f4 |
| ENCFF883FJF.tabix.bed.gz.tbi.dvc | 110.0 B | dae6c8a4cfb97eef6d56475b70f25af7 |
| genomic_resource.yaml | 2.99 KB | f6f065b0f09d55978bbb00958c385f26 |
| genomic_resource_original.yaml | 2.83 KB | 115873ae201b94f7582305596e247060 |
| statistics/ |