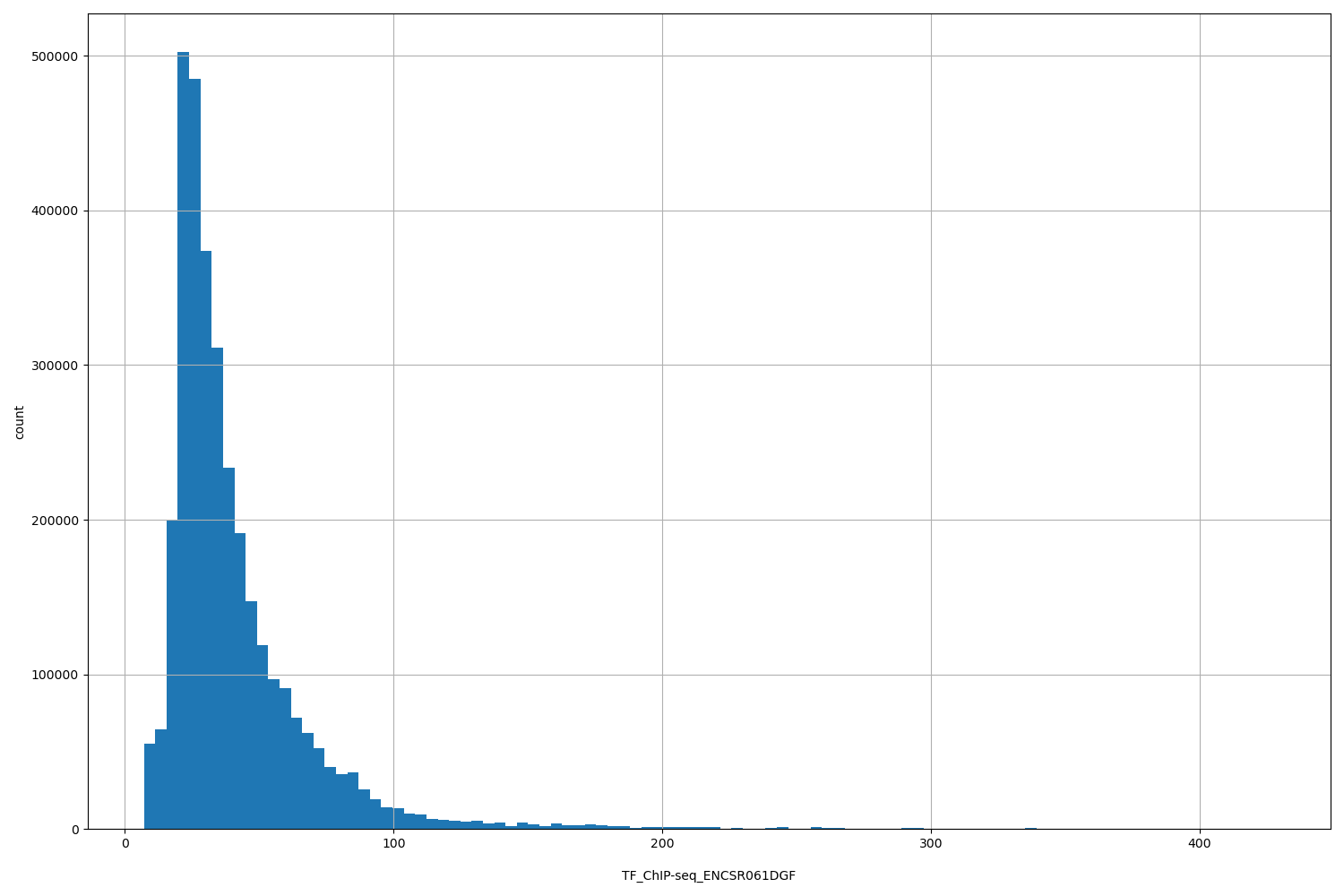
TF_ChIP-seq_ENCSR061DGF
| Id: | TF_ChIP-seq/ENCSR061DGF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR061DGF [biosamplesummary="Homo sapiens GM23338 originated from GM23248" and target="NANOG"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN497WXY|/analyses/ENCAN497WXY/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF131FBY|/files/ENCFF131FBY/}, {ENCFF509ILA|/files/ENCFF509ILA/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.67. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF735WFO|/files/ENCFF735WFO/}, {ENCFF466GNI|/files/ENCFF466GNI/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.67. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR061DGF | float |
TF_ChIP-seq_ENCSR061DGF |
TF_ChIP-seq ENCSR061DGF [biosample_summary="Homo sapiens GM23338 originated from GM23248" and target="NANOG"]
|
 |
[7.07, 428] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF735WFO.bed.gz | 196.17 KB | 29c10e45fa37cf7416d81f1e6f977d5c |
| ENCFF735WFO.bed.gz.dvc | 100.0 B | 88e7bf6bb925c127eb7013b21b33dd61 |
| ENCFF735WFO.tabix.bed.gz | 136.09 KB | 97a0da837a662dc6c4e075162c938691 |
| ENCFF735WFO.tabix.bed.gz.dvc | 106.0 B | 123f15eab5f590e7707e1ef0da2f0821 |
| ENCFF735WFO.tabix.bed.gz.tbi | 92.9 KB | 07108998b4eb8e8da91638eb6b268f8b |
| ENCFF735WFO.tabix.bed.gz.tbi.dvc | 109.0 B | f7f3143ff0748a72a698be5ba9be706d |
| genomic_resource.yaml | 2.71 KB | 48a4c214d2f27fab244a0a4cc7cf22d7 |
| genomic_resource_original.yaml | 2.59 KB | a44b699ca0a778c1d400767b718afbc7 |
| statistics/ |