TF_ChIP-seq_ENCSR000FAG
| Id: | TF_ChIP-seq/ENCSR000FAG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FAG [biosamplesummary="Homo sapiens K562" and target="MYC"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF937QDF|/files/ENCFF937QDF/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 3395233 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF788FDI|/files/ENCFF788FDI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 3192210 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN568MZG|/analyses/ENCAN568MZG/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN568MZG|/analyses/ENCAN568MZG/} has in progress subobject document {782d7403-b142-4d82-b451-531a426162d8|/documents/782d7403-b142-4d82-b451-531a426162d8/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF675PHY|/files/ENCFF675PHY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.07 and a self consistency ratio of 2.22. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
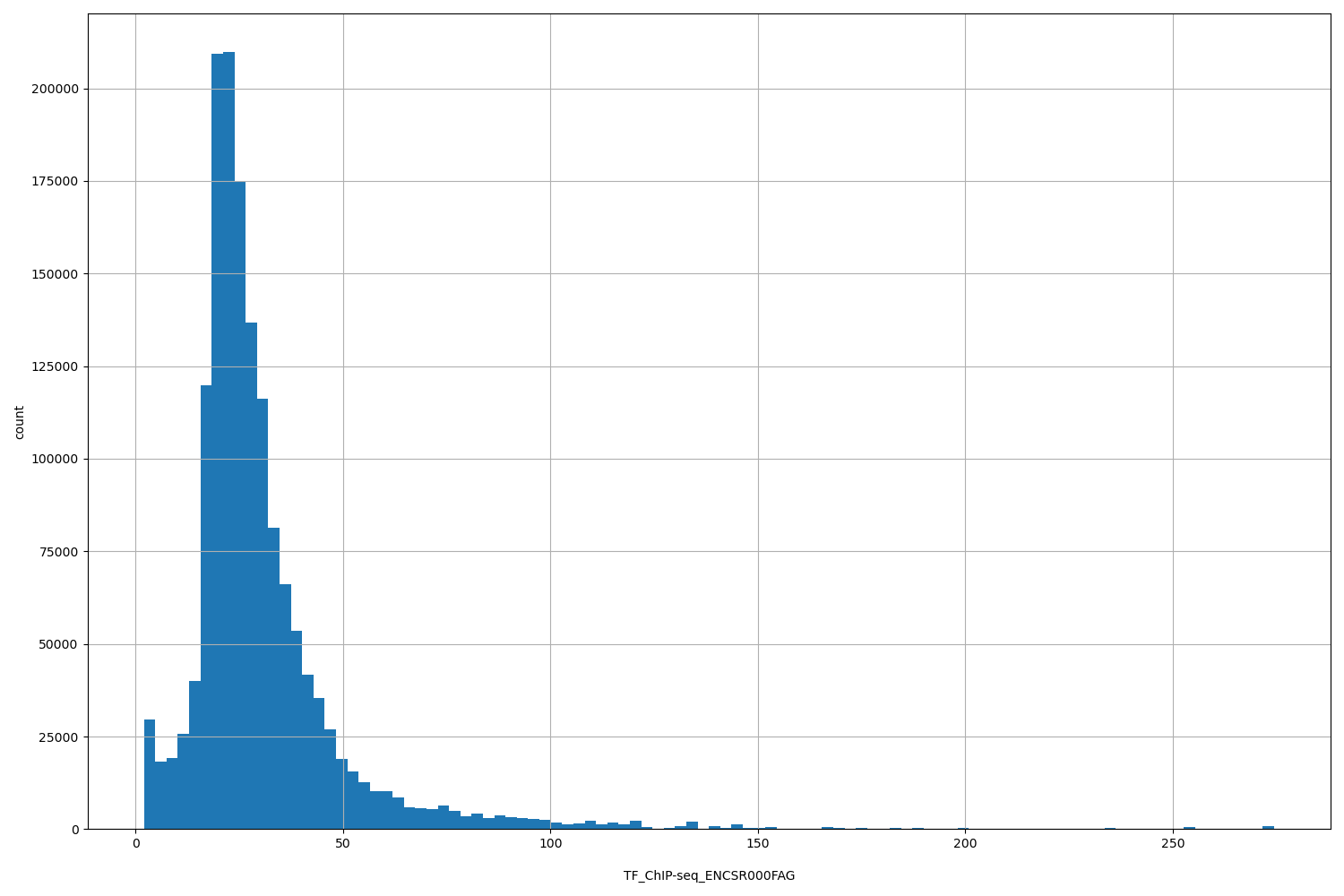
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FAG | float |
TF_ChIP-seq_ENCSR000FAG |
TF_ChIP-seq ENCSR000FAG [biosample_summary="Homo sapiens K562" and target="MYC"]
|
 |
[2, 274] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF675PHY.bed.gz | 83.46 KB | 6ac174e470b650bad989ee2c1ed328db |
| ENCFF675PHY.bed.gz.dvc | 99.0 B | 382dde5787c6b8d6a81fe28990925db8 |
| ENCFF675PHY.tabix.bed.gz | 64.36 KB | ed71c97e410b1984491f3ea1257eba6d |
| ENCFF675PHY.tabix.bed.gz.dvc | 105.0 B | abde00c230c8e7b7e086740e52f0f961 |
| ENCFF675PHY.tabix.bed.gz.tbi | 46.68 KB | becb8ef0082ef82f35d31a7c3bf8b749 |
| ENCFF675PHY.tabix.bed.gz.tbi.dvc | 109.0 B | eefcc5395c63cac0d3e8e74695c8c977 |
| genomic_resource.yaml | 3.09 KB | a94acbd912c64deaa25a610ae5c77515 |
| genomic_resource_original.yaml | 3.0 KB | 534ec1511305efd6a3f4d1d41c41ed00 |
| statistics/ |