TF_ChIP-seq_ENCSR000EYZ
| Id: | TF_ChIP-seq/ENCSR000EYZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EYZ [biosamplesummary="Homo sapiens GM12878" and target="FOS"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF139BSO|/files/ENCFF139BSO/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 1995810 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting FOS-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF814AEX|/files/ENCFF814AEX/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 2973143 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting FOS-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF940GUI|/files/ENCFF940GUI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 2901575 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting FOS-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN950YYJ|/analyses/ENCAN950YYJ/} has in progress subobject document {706a4d18-ce66-462c-a7b4-e28dead55d36|/documents/706a4d18-ce66-462c-a7b4-e28dead55d36/} audit_internal_action: Released analysis {ENCAN950YYJ|/analyses/ENCAN950YYJ/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF571DGT|/files/ENCFF571DGT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.63 and a self consistency ratio of 5.53. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
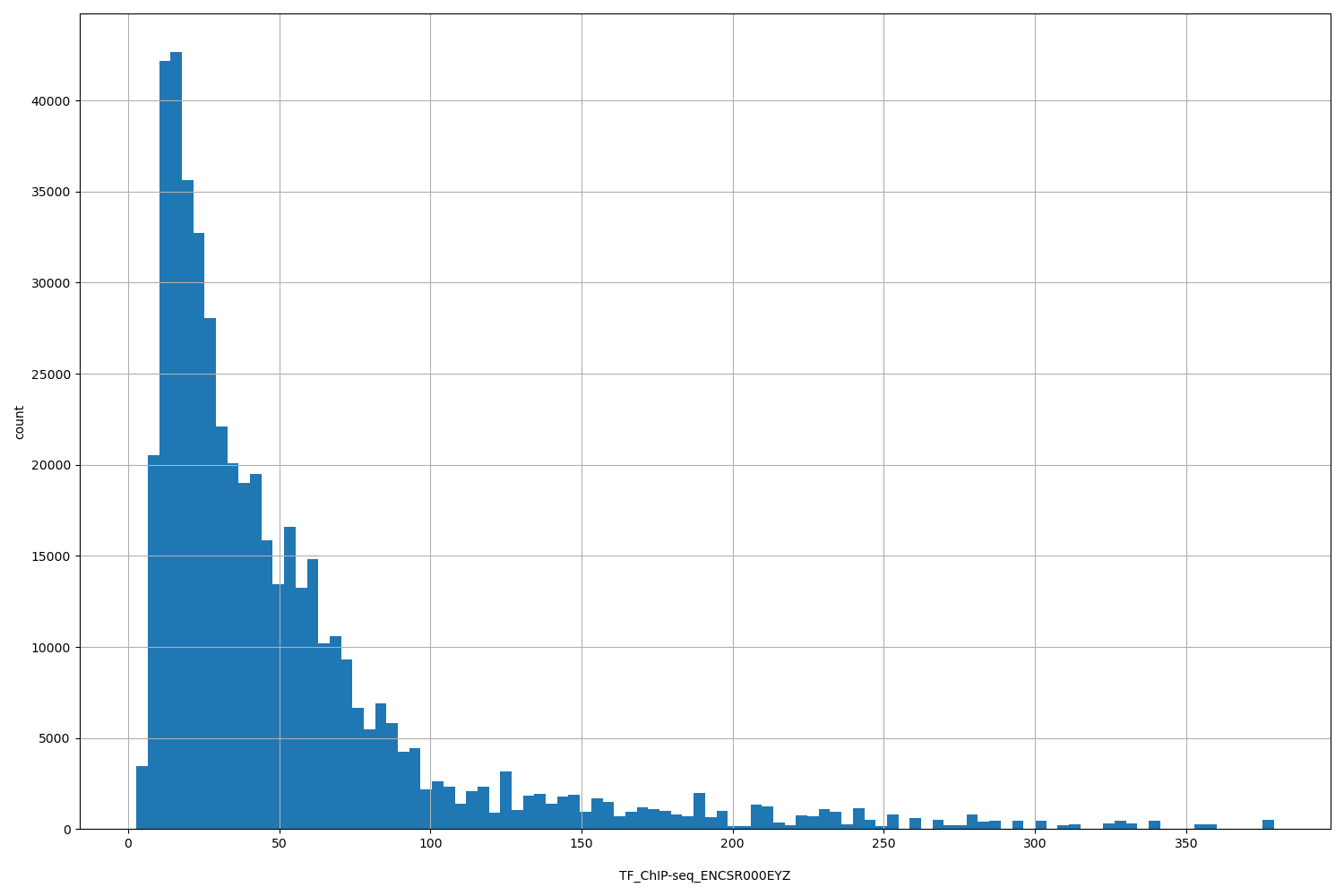
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EYZ | float |
TF_ChIP-seq_ENCSR000EYZ |
TF_ChIP-seq ENCSR000EYZ [biosample_summary="Homo sapiens GM12878" and target="FOS"]
|
 |
[2.71, 379] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF571DGT.bed.gz | 34.97 KB | 7936abaa01edc683810b9a4de2d78900 |
| ENCFF571DGT.bed.gz.dvc | 99.0 B | c9653d309caa4c815d7363657852dd3b |
| ENCFF571DGT.tabix.bed.gz | 25.09 KB | ac718317cb0dc8d849a5bdee4e34a4fe |
| ENCFF571DGT.tabix.bed.gz.dvc | 105.0 B | 88e135ac34d7c9dea3aa76fd127a2b1e |
| ENCFF571DGT.tabix.bed.gz.tbi | 24.67 KB | 369891ef5b9062fe816f97786147f604 |
| ENCFF571DGT.tabix.bed.gz.tbi.dvc | 109.0 B | 2f5f5ba277fc66d5df74df410e5f682a |
| genomic_resource.yaml | 3.6 KB | bdfd4dd68f0eef454299af20ccb92437 |
| genomic_resource_original.yaml | 3.51 KB | e5f0b62e2930b631e40e94cc01f8e1e3 |
| statistics/ |