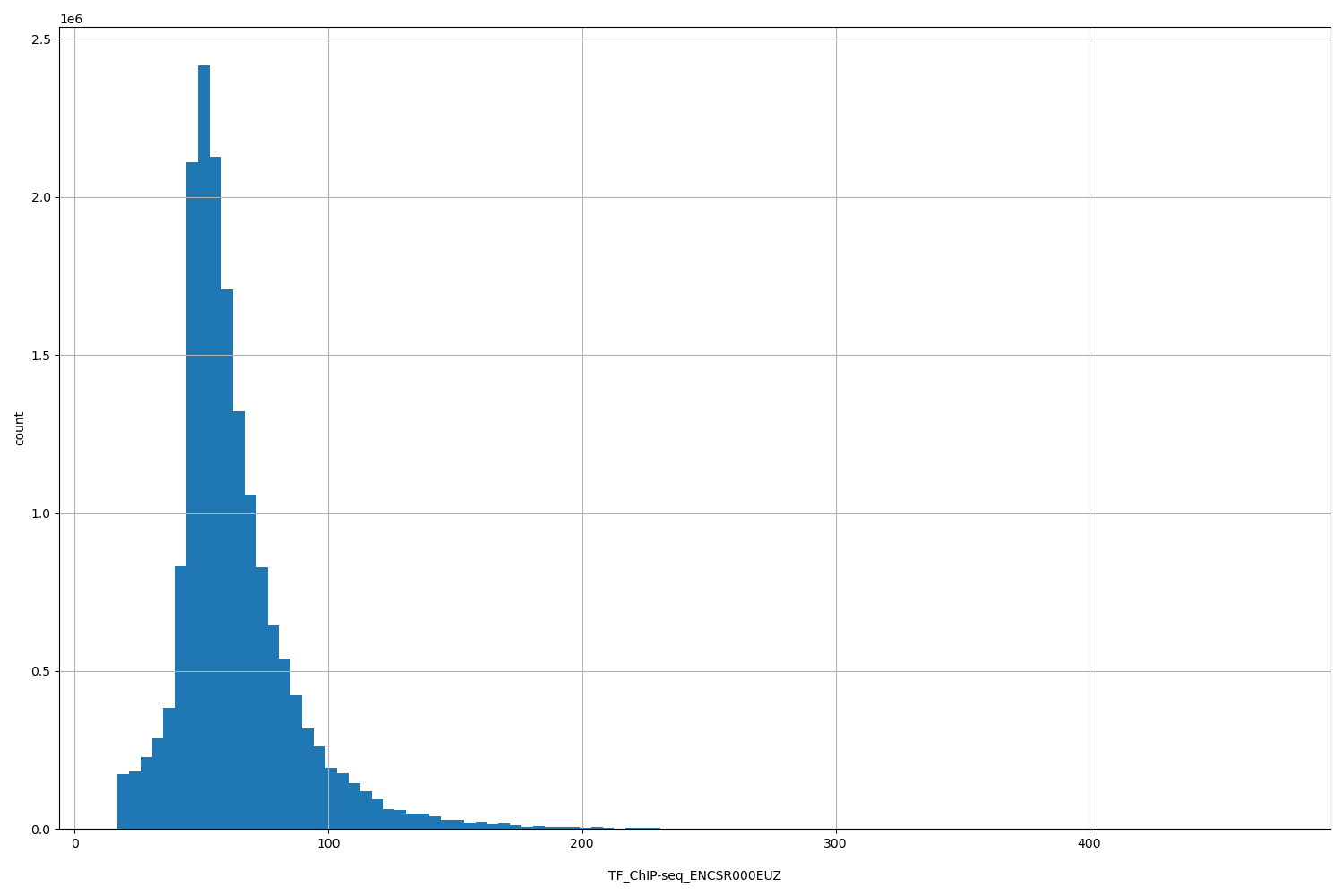
TF_ChIP-seq_ENCSR000EUZ
| Id: | TF_ChIP-seq/ENCSR000EUZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EUZ [biosamplesummary="Homo sapiens HEK293" and target="TRIM28"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN593AVV|/analyses/ENCAN593AVV/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF642DXY|/files/ENCFF642DXY/}, {ENCFF721FYE|/files/ENCFF721FYE/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.80 and a self consistency ratio of 1.30. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF046VYV|/files/ENCFF046VYV/}, {ENCFF210BOH|/files/ENCFF210BOH/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.80 and a self consistency ratio of 1.30. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EUZ | float |
TF_ChIP-seq_ENCSR000EUZ |
TF_ChIP-seq ENCSR000EUZ [biosample_summary="Homo sapiens HEK293" and target="TRIM28"]
|
 |
[16.8, 472] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF721FYE.bed.gz | 533.76 KB | e683f117e45e4e61bdd765e45e363858 |
| ENCFF721FYE.bed.gz.dvc | 100.0 B | 0735d270d952e17e0484f00fccd19626 |
| ENCFF721FYE.tabix.bed.gz | 416.23 KB | 3dbee4939d11f73b6767aff16ac1f91b |
| ENCFF721FYE.tabix.bed.gz.dvc | 106.0 B | 47fe3d028b9df185f1251b0315c2111a |
| ENCFF721FYE.tabix.bed.gz.tbi | 170.51 KB | 65736b4ff49bb0ca76582998634da238 |
| ENCFF721FYE.tabix.bed.gz.tbi.dvc | 110.0 B | 9092cb1aef7050e33c625981ce1d8d5d |
| genomic_resource.yaml | 2.7 KB | 9d4abf679109e1eb3aa2860e22ce8b12 |
| genomic_resource_original.yaml | 2.6 KB | a50d2cd74e3e162bd4aea12846612154 |
| statistics/ |