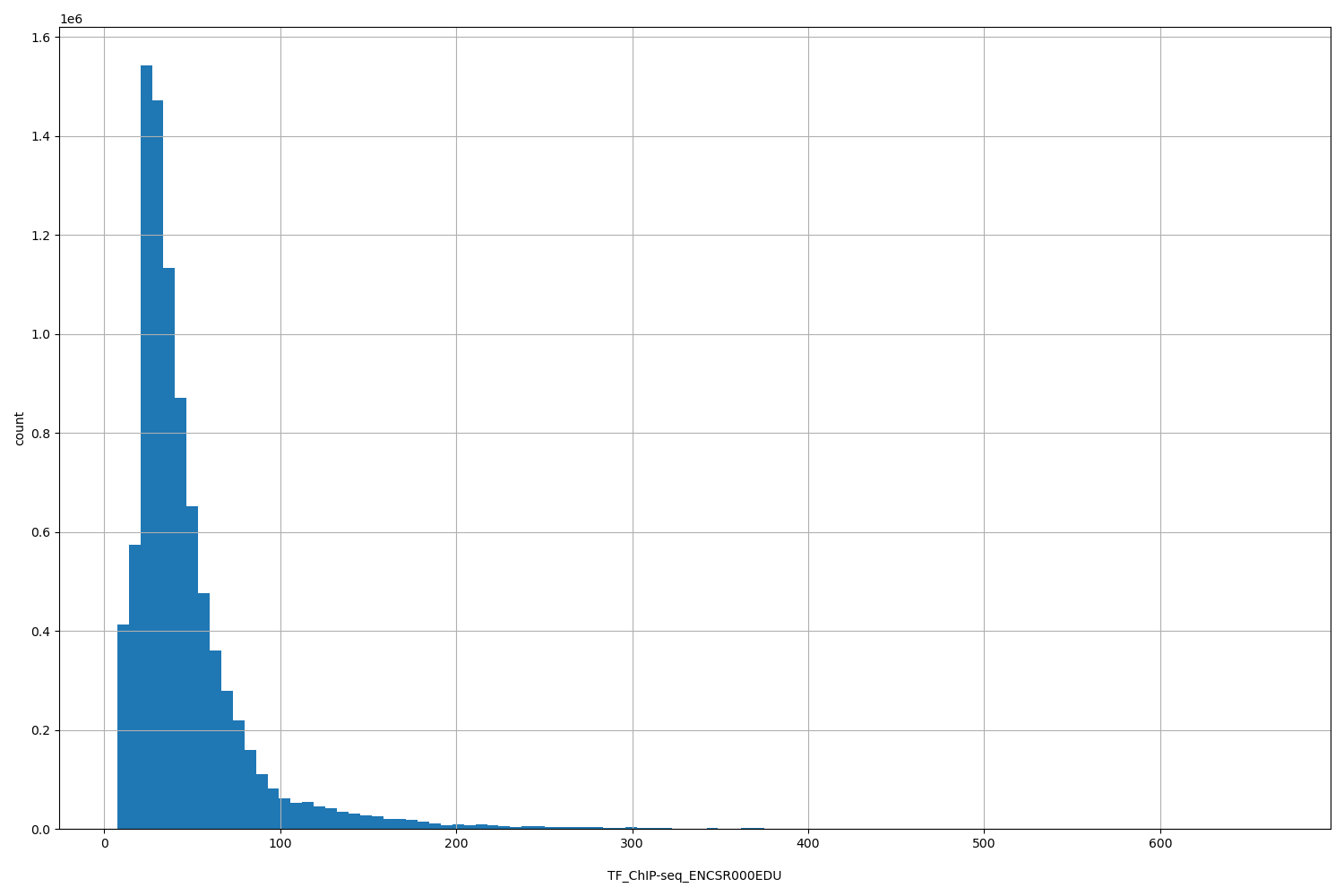
TF_ChIP-seq_ENCSR000EDU
| Id: | TF_ChIP-seq/ENCSR000EDU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDU [biosamplesummary="Homo sapiens HepG2" and target="MXI1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN629KHG|/analyses/ENCAN629KHG/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN629KHG|/analyses/ENCAN629KHG/} has in progress subobject document {b9ed06bc-2e72-46e1-b318-0bba344a8665|/documents/b9ed06bc-2e72-46e1-b318-0bba344a8665/} audit_warning: Processed alignments file {ENCFF838XHB|/files/ENCFF838XHB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14072850 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MXI1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF153QSO|/files/ENCFF153QSO/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19476019 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MXI1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF599KZZ|/files/ENCFF599KZZ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.56 and a self consistency ratio of 13.70. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDU | float |
TF_ChIP-seq_ENCSR000EDU |
TF_ChIP-seq ENCSR000EDU [biosample_summary="Homo sapiens HepG2" and target="MXI1"]
|
 |
[7.45, 664] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF599KZZ.bed.gz | 420.11 KB | 89970f3a10119bc4f03752d80b1d7c81 |
| ENCFF599KZZ.bed.gz.dvc | 100.0 B | 6c4690f772c064a698c192303b63a261 |
| ENCFF599KZZ.tabix.bed.gz | 326.53 KB | 917833dcc5401c8d4ed91566ec78ceb0 |
| ENCFF599KZZ.tabix.bed.gz.dvc | 106.0 B | d853ea2e233ec2f0cef8152b434dc3d3 |
| ENCFF599KZZ.tabix.bed.gz.tbi | 138.07 KB | a9fb0516db8171bce230b67ca560e8c7 |
| ENCFF599KZZ.tabix.bed.gz.tbi.dvc | 110.0 B | c0b99a1674f66d96bbbcc490bb845d00 |
| genomic_resource.yaml | 3.1 KB | 66d2bc4627ea663081b31e1e7fb7868b |
| genomic_resource_original.yaml | 3.01 KB | 76692977195792a17d71a15e9118c402 |
| statistics/ |