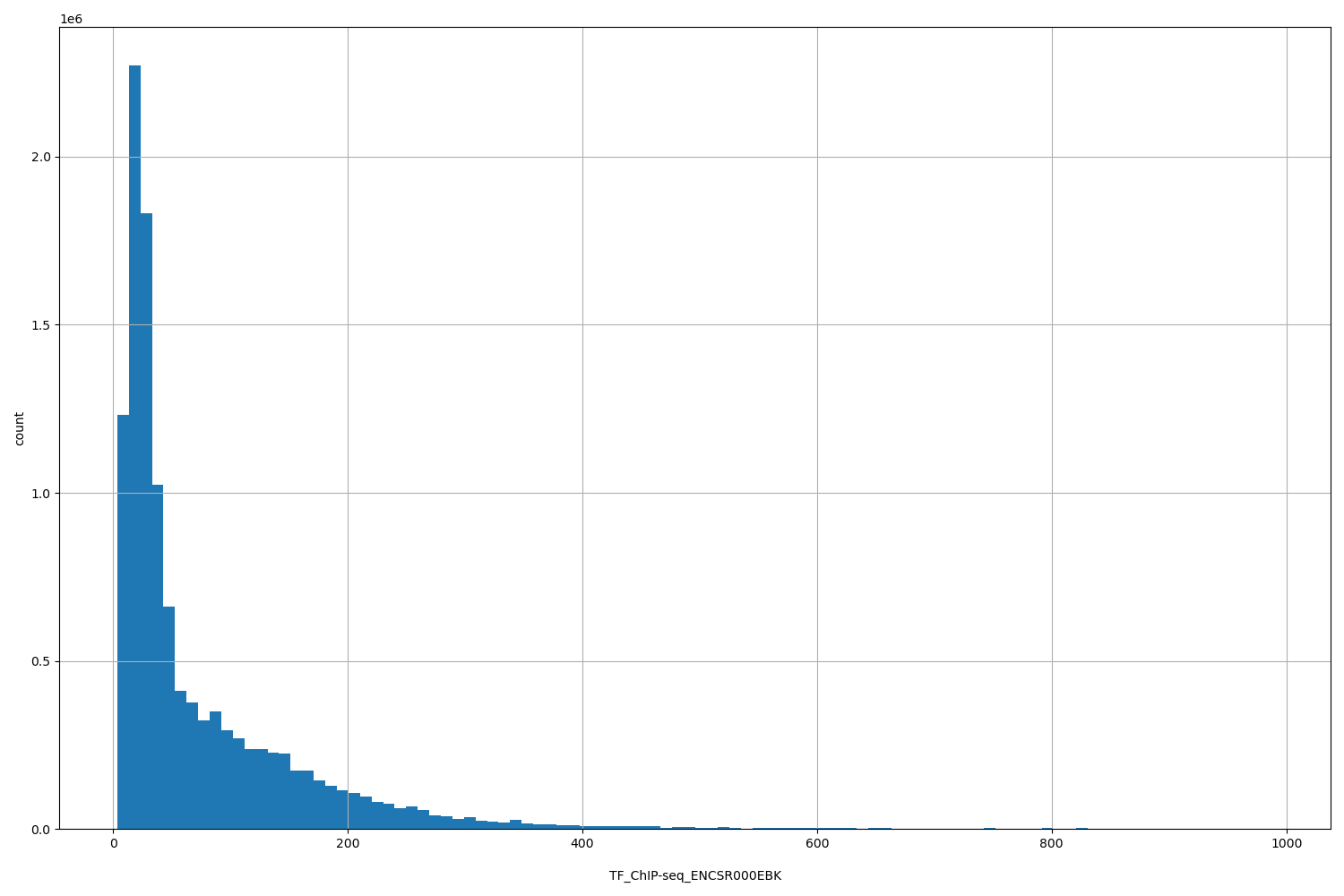
TF_ChIP-seq_ENCSR000EBK
| Id: | TF_ChIP-seq/ENCSR000EBK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EBK [biosamplesummary="Homo sapiens GM19193" and target="POLR2A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3, Rep 4 summary: output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF660FZH|/files/ENCFF660FZH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 2857272 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF835RFW|/files/ENCFF835RFW/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 4556545 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF169HFV|/files/ENCFF169HFV/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 3626200 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN286CLG|/analyses/ENCAN286CLG/} has in progress subobject document {a4cdfb8a-af24-4c92-8fd3-6071701f4564|/documents/a4cdfb8a-af24-4c92-8fd3-6071701f4564/} audit_internal_action: Released analysis {ENCAN286CLG|/analyses/ENCAN286CLG/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF071IGH|/files/ENCFF071IGH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 5906971 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EBK | float |
TF_ChIP-seq_ENCSR000EBK |
TF_ChIP-seq ENCSR000EBK [biosample_summary="Homo sapiens GM19193" and target="POLR2A"]
|
 |
[3.57, 988] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF519DDH.bed.gz | 521.19 KB | f73575b9c1cd841d5a770a0a55f8bafd |
| ENCFF519DDH.bed.gz.dvc | 100.0 B | b3461e6476f9db7abfa5e14470fddc7c |
| ENCFF519DDH.tabix.bed.gz | 379.58 KB | 0b063d4a7c1827fe20f8cb37bdbd95d1 |
| ENCFF519DDH.tabix.bed.gz.dvc | 106.0 B | c4ad229d13ead070e58a4ad0472bbae9 |
| ENCFF519DDH.tabix.bed.gz.tbi | 126.05 KB | e2f183e9174683cd31de1083ac1bc403 |
| ENCFF519DDH.tabix.bed.gz.tbi.dvc | 110.0 B | 1a4e79623303813292e1286f5f724781 |
| genomic_resource.yaml | 3.59 KB | ede974f8f77206bb64bcb2191ab31646 |
| genomic_resource_original.yaml | 3.49 KB | f0f2d9e4ed3e68aae8f9457480101bc5 |
| statistics/ |