TF_ChIP-seq_ENCSR000DRE
| Id: | TF_ChIP-seq/ENCSR000DRE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DRE [biosamplesummary="Homo sapiens GM12865" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 2, Rep 3 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN846DYX|/analyses/ENCAN846DYX/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF965YZI|/files/ENCFF965YZI/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_not_compliant: Processed alignments file {ENCFF383IOB|/files/ENCFF383IOB/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 8105817 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF655BNX|/files/ENCFF655BNX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 13550892 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
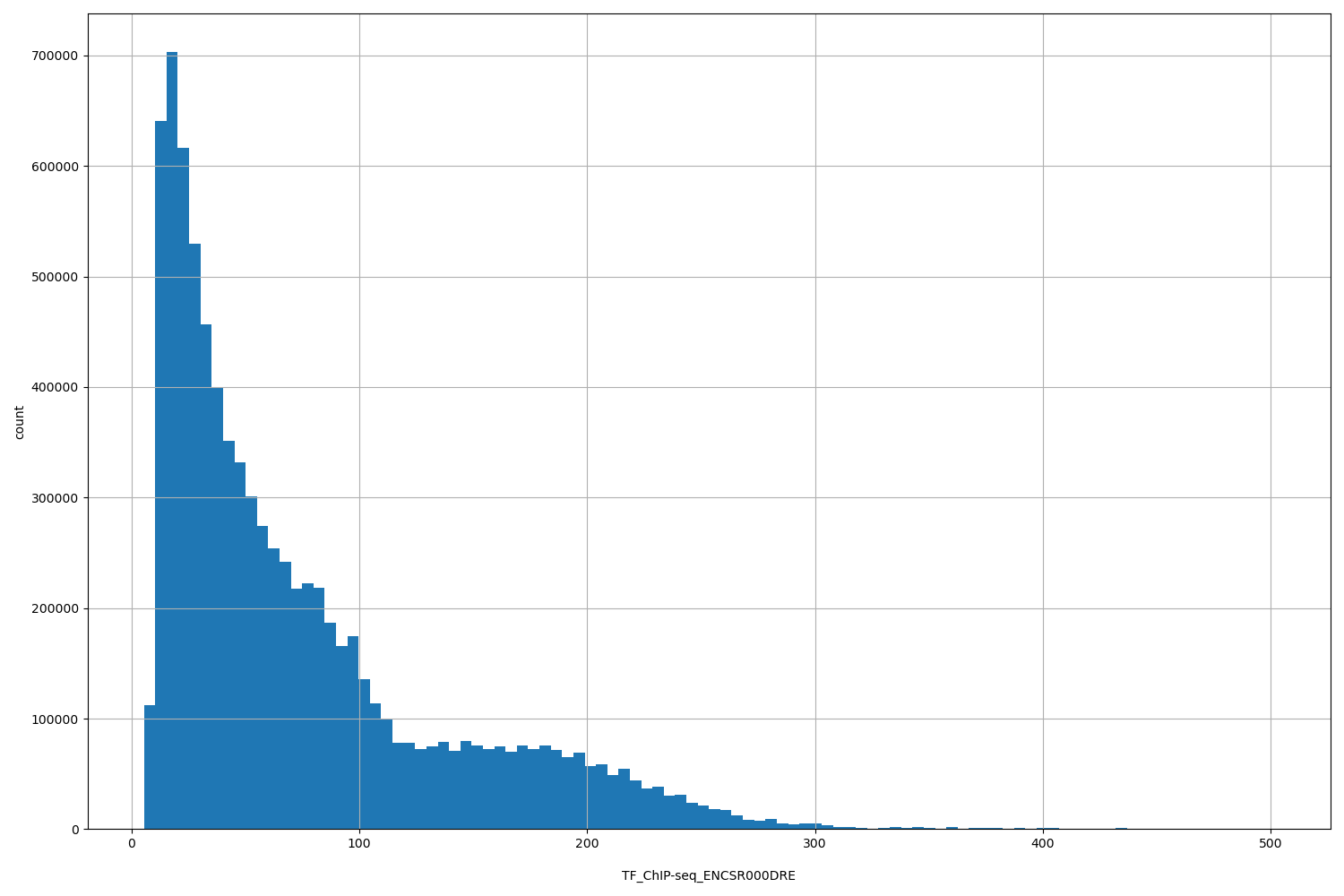
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DRE | float |
TF_ChIP-seq_ENCSR000DRE |
TF_ChIP-seq ENCSR000DRE [biosample_summary="Homo sapiens GM12865" and target="CTCF"]
|
 |
[5.41, 501] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF965YZI.bed.gz | 616.05 KB | 941a8a409acd3664e72b358da282ea06 |
| ENCFF965YZI.bed.gz.dvc | 100.0 B | 597869a4655c9f31f3a47ac633368737 |
| ENCFF965YZI.tabix.bed.gz | 458.36 KB | abcd9501d30dfd39586e9e70497bc2f6 |
| ENCFF965YZI.tabix.bed.gz.dvc | 106.0 B | 9252a76badc70f3b3c5e04057ffb7a82 |
| ENCFF965YZI.tabix.bed.gz.tbi | 258.43 KB | 874a5f135038bb63dd30163bd4d7fd39 |
| ENCFF965YZI.tabix.bed.gz.tbi.dvc | 110.0 B | 02436b64730bf3a65f4003dd6110832e |
| genomic_resource.yaml | 2.92 KB | 308855cbc100489b003aebd00f1eeab4 |
| genomic_resource_original.yaml | 2.82 KB | e80040a687321f7a9914962e71590470 |
| statistics/ |