TF_ChIP-seq_ENCSR000DOK
| Id: | TF_ChIP-seq/ENCSR000DOK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DOK [biosamplesummary="Homo sapiens K562" and target="BDP1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN438QXX|/analyses/ENCAN438QXX/} has in progress subobject document {797f5e55-5642-41b7-80ad-a5d1d788897c|/documents/797f5e55-5642-41b7-80ad-a5d1d788897c/} audit_internal_action: Released analysis {ENCAN438QXX|/analyses/ENCAN438QXX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF039NAG|/files/ENCFF039NAG/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6091980 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BDP1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF264LHV|/files/ENCFF264LHV/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6082982 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BDP1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF938MMD|/files/ENCFF938MMD/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6076335 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BDP1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
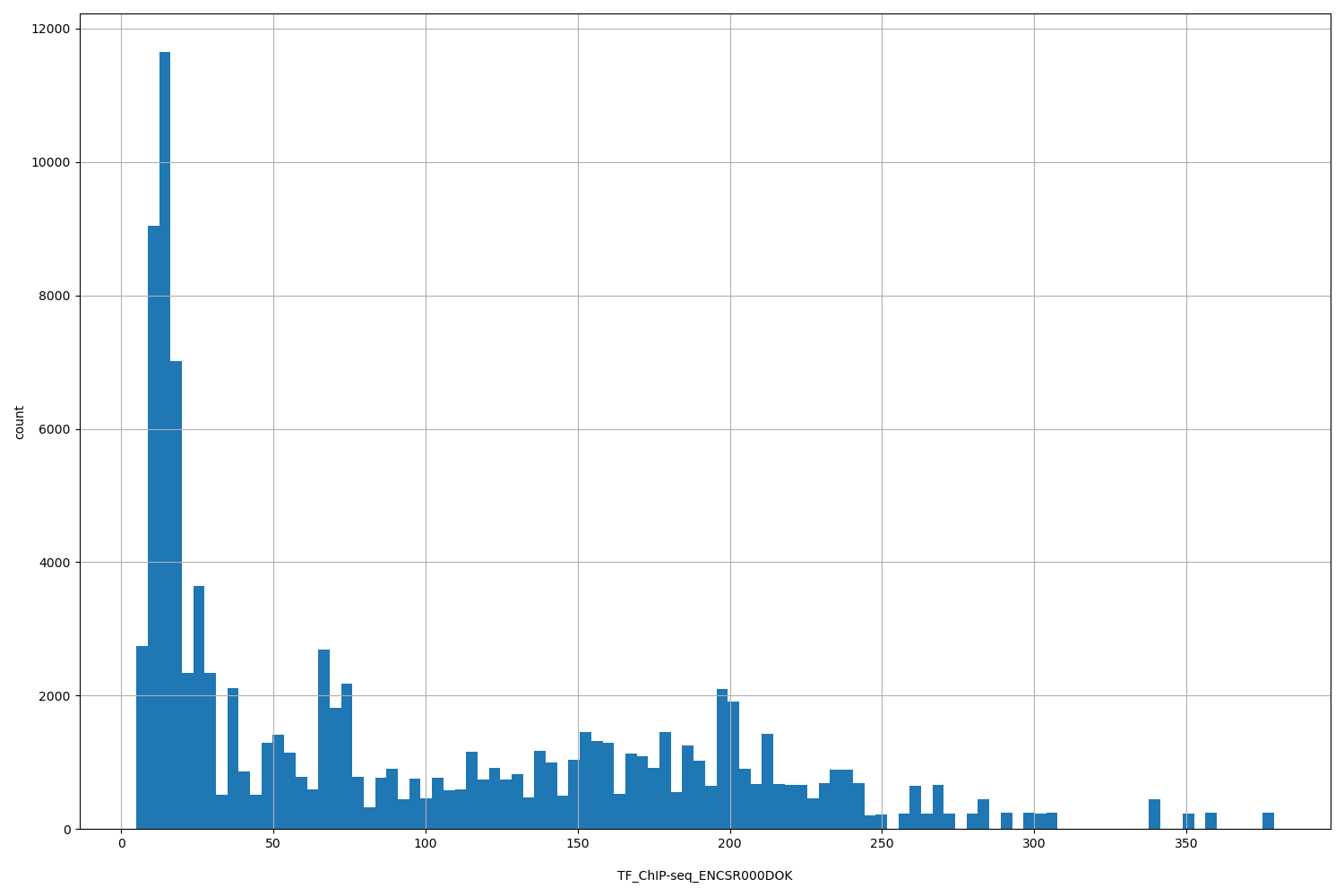
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DOK | float |
TF_ChIP-seq_ENCSR000DOK |
TF_ChIP-seq ENCSR000DOK [biosample_summary="Homo sapiens K562" and target="BDP1"]
|
 |
[4.91, 379] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF284FTY.bed.gz | 8.24 KB | 2eb136799de1f5ffdab8a23d4bd4da74 |
| ENCFF284FTY.bed.gz.dvc | 98.0 B | 5acb480e1013b61789bb3e4eb76b1546 |
| ENCFF284FTY.tabix.bed.gz | 5.62 KB | cb61c6c86b7caf0c269ce25a46fbed9c |
| ENCFF284FTY.tabix.bed.gz.dvc | 104.0 B | ba40926be3db8588ac59c2d12494cc47 |
| ENCFF284FTY.tabix.bed.gz.tbi | 6.94 KB | 3cb41fd70e007563ffa1c495f8ce0e03 |
| ENCFF284FTY.tabix.bed.gz.tbi.dvc | 108.0 B | 19dac96b446491e185d4263db2034ebe |
| genomic_resource.yaml | 3.08 KB | 3248ea7cc9b926d7f3d956f0f47dac87 |
| genomic_resource_original.yaml | 2.99 KB | 1f2a6563601784a83e93de6d30faf6f5 |
| statistics/ |