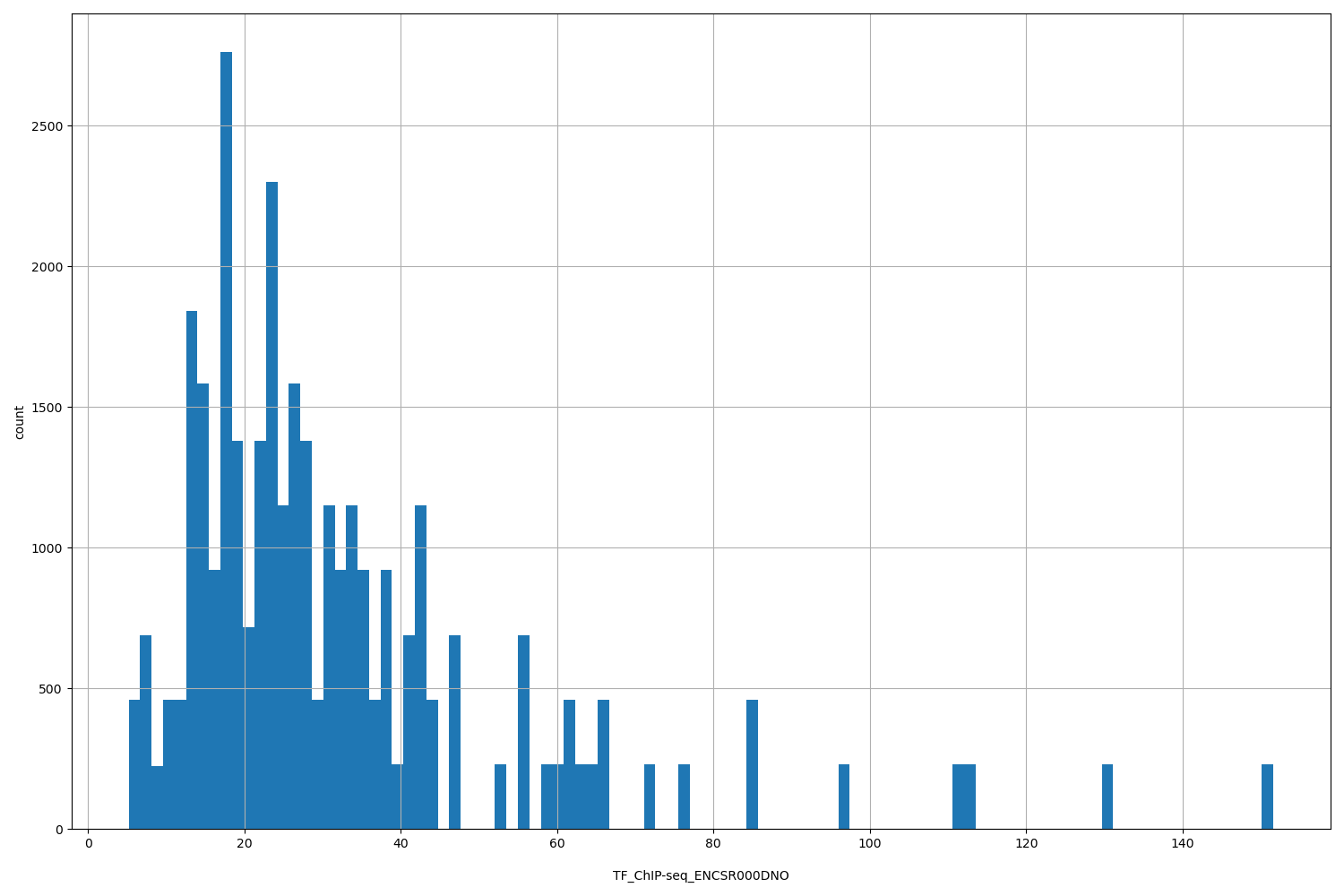
TF_ChIP-seq_ENCSR000DNO
| Id: | TF_ChIP-seq/ENCSR000DNO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DNO [biosamplesummary="Homo sapiens GM12878" and target="KAT2A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: conservative IDR thresholded peaks audit_internal_action: Released analysis {ENCAN560LJC|/analyses/ENCAN560LJC/} has in progress subobject document {36ac3c42-d441-4926-a7b1-f9b1cdf33cb2|/documents/36ac3c42-d441-4926-a7b1-f9b1cdf33cb2/} audit_internal_action: Released analysis {ENCAN560LJC|/analyses/ENCAN560LJC/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF495LKP|/files/ENCFF495LKP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13350061 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting KAT2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF117AZM|/files/ENCFF117AZM/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13381185 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting KAT2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DNO | float |
TF_ChIP-seq_ENCSR000DNO |
TF_ChIP-seq ENCSR000DNO [biosample_summary="Homo sapiens GM12878" and target="KAT2A"]
|
 |
[5.22, 152] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF333PDI.bed.gz | 2.74 KB | 8e24be37bdeb76eea8deb3cb38013683 |
| ENCFF333PDI.bed.gz.dvc | 98.0 B | 74cd7814d8908a88e1dd35e36104865e |
| ENCFF333PDI.tabix.bed.gz | 1.79 KB | 898c5716d2549f4f10901ac23b8d8190 |
| ENCFF333PDI.tabix.bed.gz.dvc | 104.0 B | f2971062569bacd2004ce44b3df58f46 |
| ENCFF333PDI.tabix.bed.gz.tbi | 4.21 KB | 5634af5a2fd803ea0f3abd57b7d51e03 |
| ENCFF333PDI.tabix.bed.gz.tbi.dvc | 108.0 B | ed5108fe2b3d79b2f9e637b027a68920 |
| genomic_resource.yaml | 2.72 KB | 9a95409b0059d032a9f8b5db0edba3c6 |
| genomic_resource_original.yaml | 2.62 KB | 2dc5757ab06f54c59c18382a9be84422 |
| statistics/ |