TF_ChIP-seq_ENCSR000DLW
| Id: | TF_ChIP-seq/ENCSR000DLW |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DLW [biosamplesummary="Homo sapiens endothelial cell of umbilical vein newborn" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: newborn output_type: conservative IDR thresholded peaks audit_internal_action: Released analysis {ENCAN996IYY|/analyses/ENCAN996IYY/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN996IYY|/analyses/ENCAN996IYY/} has in progress subobject document {2c6beb16-84f5-4228-bcb7-9808a4d9e97d|/documents/2c6beb16-84f5-4228-bcb7-9808a4d9e97d/} audit_not_compliant: Processed alignments file {ENCFF690KEO|/files/ENCFF690KEO/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 7840777 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF307WMW|/files/ENCFF307WMW/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 8750997 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
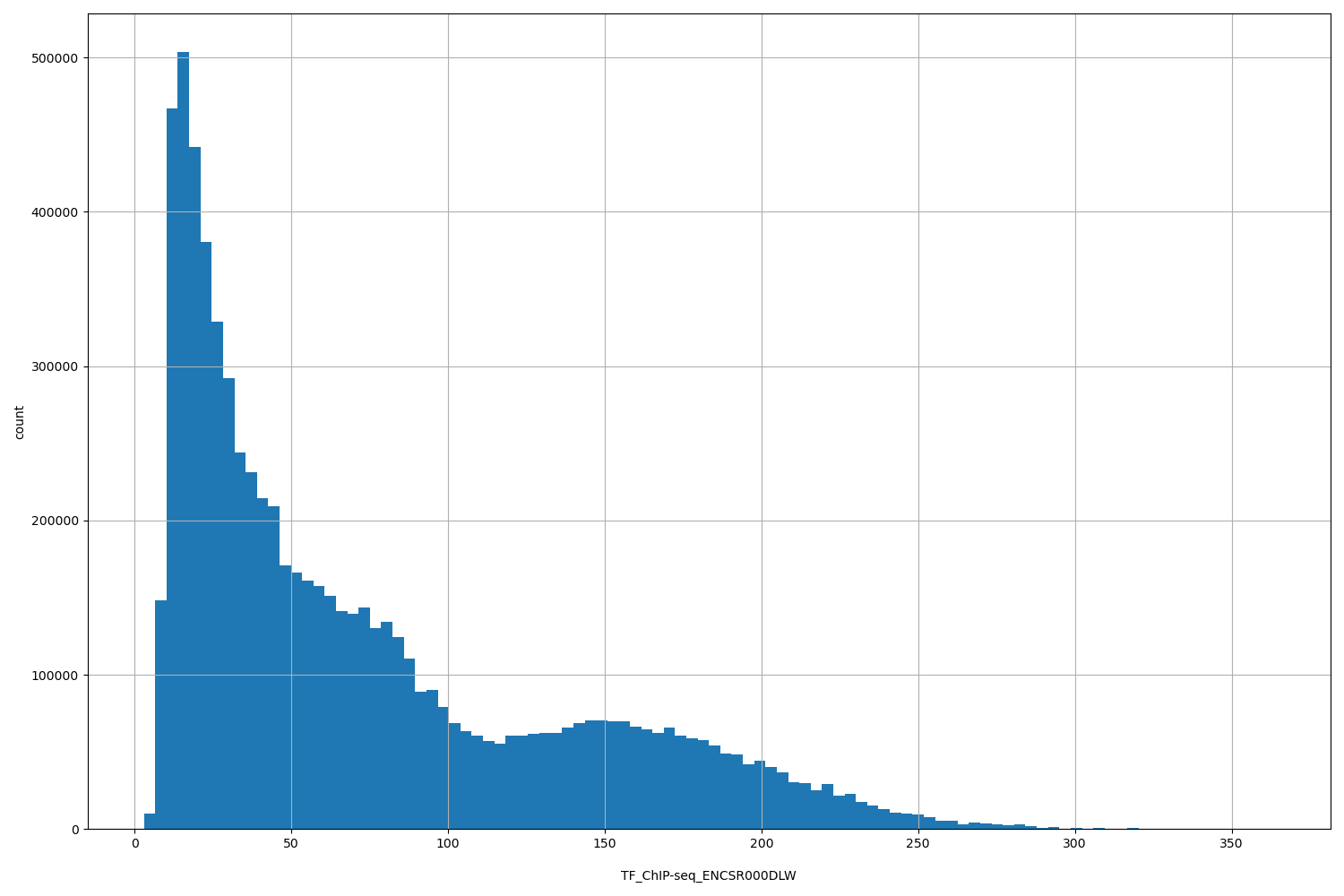
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DLW | float |
TF_ChIP-seq_ENCSR000DLW |
TF_ChIP-seq ENCSR000DLW [biosample_summary="Homo sapiens endothelial cell of umbilical vein newborn" and target="CTCF"]
|
 |
[2.99, 364] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF664CQH.bed.gz | 684.62 KB | 816fe007ff04efee85a2854e12261363 |
| ENCFF664CQH.bed.gz.dvc | 100.0 B | a3c6ae8001fb9de13720ca6a516b4667 |
| ENCFF664CQH.tabix.bed.gz | 531.88 KB | 81a5a12d6fac658bea7dfda4c11d1071 |
| ENCFF664CQH.tabix.bed.gz.dvc | 106.0 B | e7307936a33ec151d23309e2ecf79f38 |
| ENCFF664CQH.tabix.bed.gz.tbi | 289.71 KB | e1342ea4b694618a5e68832dcb72ea73 |
| ENCFF664CQH.tabix.bed.gz.tbi.dvc | 110.0 B | 5aa7ffcecf67c26eaf71210d566df2e0 |
| genomic_resource.yaml | 2.81 KB | 8f3be01bd712225aec6a39e8c1b32468 |
| genomic_resource_original.yaml | 2.69 KB | 5ca29b0406a89fc11f0bc4539f4f29bc |
| statistics/ |