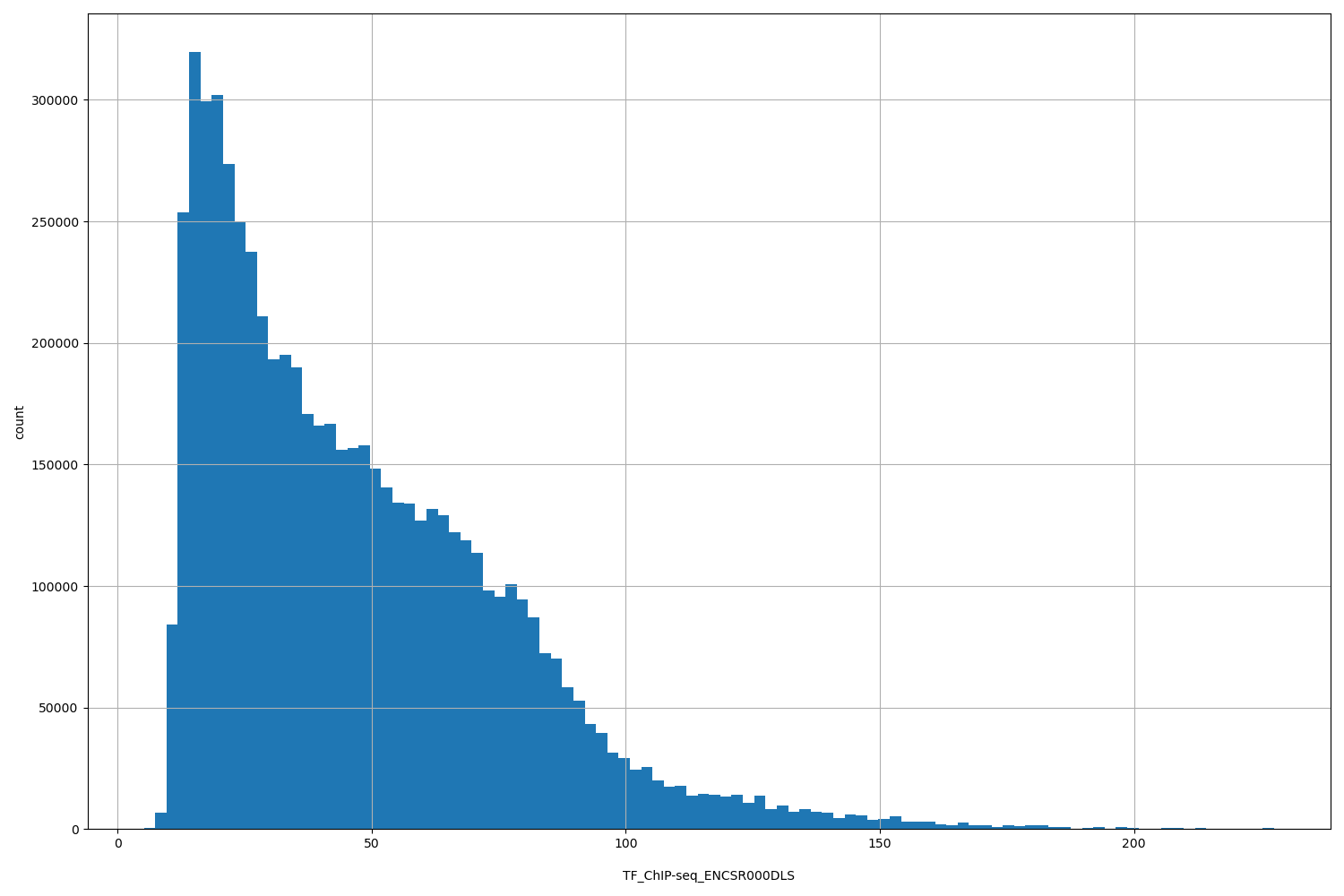
TF_ChIP-seq_ENCSR000DLS
| Id: | TF_ChIP-seq/ENCSR000DLS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DLS [biosamplesummary="Homo sapiens HepG2" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF255JDC|/files/ENCFF255JDC/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 4888689 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN700ZXA|/analyses/ENCAN700ZXA/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN700ZXA|/analyses/ENCAN700ZXA/} has in progress subobject document {79342162-8cb4-472a-a5ee-59e356f7ef31|/documents/79342162-8cb4-472a-a5ee-59e356f7ef31/} audit_not_compliant: Processed alignments file {ENCFF485DWL|/files/ENCFF485DWL/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 5054757 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DLS | float |
TF_ChIP-seq_ENCSR000DLS |
TF_ChIP-seq ENCSR000DLS [biosample_summary="Homo sapiens HepG2" and target="CTCF"]
|
 |
[5.18, 228] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF240MUS.bed.gz | 645.38 KB | bf52a338189889328b60978d375ec94c |
| ENCFF240MUS.bed.gz.dvc | 100.0 B | 6b247a73a72ed9260591ed8f27ac4fc1 |
| ENCFF240MUS.tabix.bed.gz | 514.87 KB | ae5ed3894f81cbc86845c8ae8367e6ae |
| ENCFF240MUS.tabix.bed.gz.dvc | 106.0 B | 73a91d0347044a9a6ab3d58917e3e4bd |
| ENCFF240MUS.tabix.bed.gz.tbi | 288.57 KB | 72e24148bb2b8ebec6f7f981655a25f5 |
| ENCFF240MUS.tabix.bed.gz.tbi.dvc | 110.0 B | b8570f90fb5d067b2504c6dd2f711d2d |
| genomic_resource.yaml | 2.56 KB | 9f24ce0bd18ab6f40e3caea9632232f9 |
| genomic_resource_original.yaml | 2.47 KB | 3513e6910f68087d35c5083b63a7a182 |
| statistics/ |