TF_ChIP-seq_ENCSR000BVC
| Id: | TF_ChIP-seq/ENCSR000BVC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BVC [biosamplesummary="Homo sapiens SK-N-SH" and target="MEF2A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN914ZWB|/analyses/ENCAN914ZWB/} has in progress subobject document {0d42a823-c17d-4c79-a848-9ee4ad30851d|/documents/0d42a823-c17d-4c79-a848-9ee4ad30851d/} audit_internal_action: Released analysis {ENCAN914ZWB|/analyses/ENCAN914ZWB/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF808BAL|/files/ENCFF808BAL/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline has 13381683 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MEF2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF146AYQ|/files/ENCFF146AYQ/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.90. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF053MLP|/files/ENCFF053MLP/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.90. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
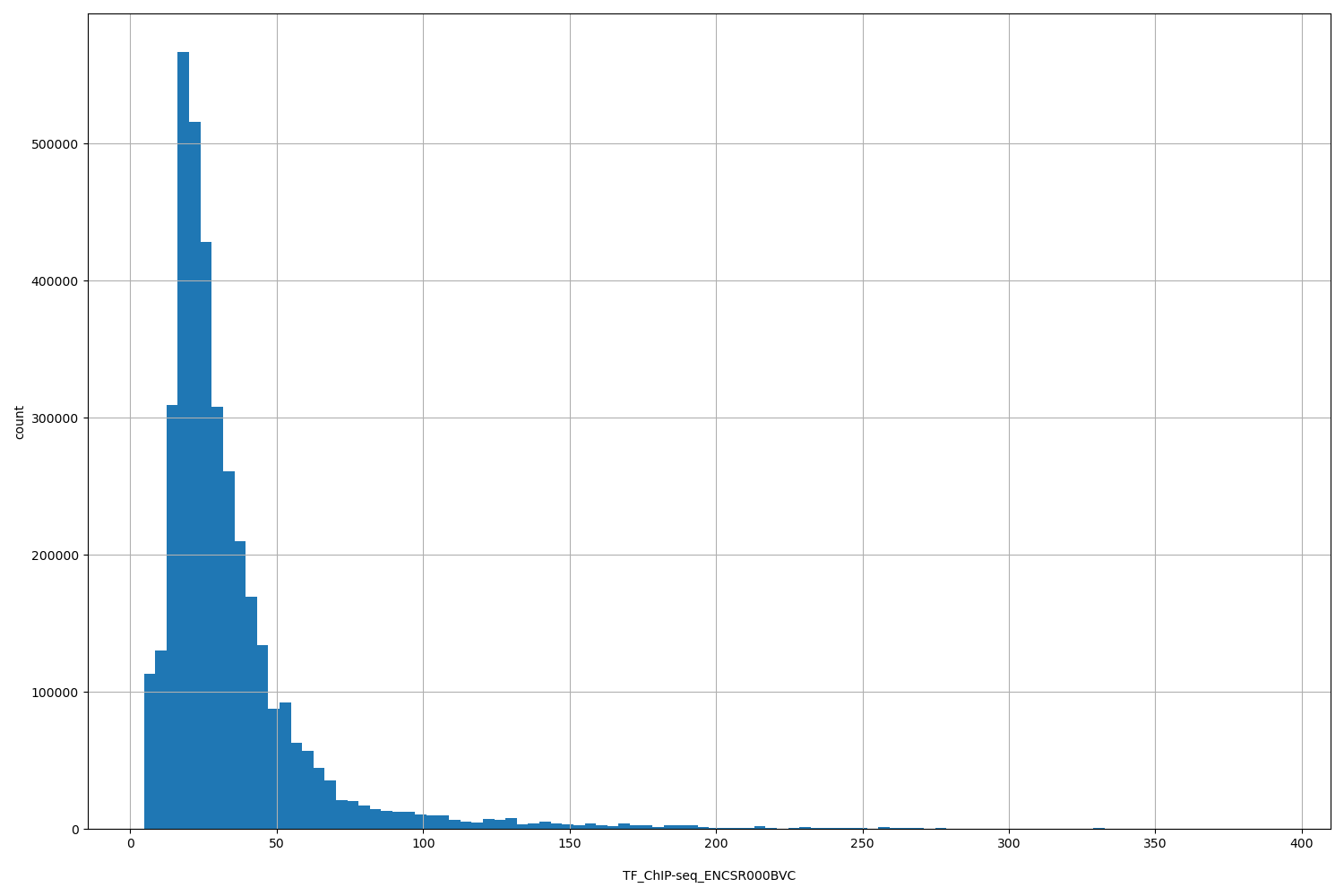
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BVC | float |
TF_ChIP-seq_ENCSR000BVC |
TF_ChIP-seq ENCSR000BVC [biosample_summary="Homo sapiens SK-N-SH" and target="MEF2A"]
|
 |
[4.64, 391] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF146AYQ.bed.gz | 203.9 KB | 97e9c54526a1d5520ec3af0658a7f17b |
| ENCFF146AYQ.bed.gz.dvc | 100.0 B | cea23040b6ac95af0b6952d25402a2a0 |
| ENCFF146AYQ.tabix.bed.gz | 145.42 KB | 9a7a13c8bf175297c87f98a74fda00d0 |
| ENCFF146AYQ.tabix.bed.gz.dvc | 106.0 B | f9233b85bdd0855f7bcc6b1d3a6a770e |
| ENCFF146AYQ.tabix.bed.gz.tbi | 85.57 KB | d2401633b8b606f409850654131976f2 |
| ENCFF146AYQ.tabix.bed.gz.tbi.dvc | 109.0 B | ec24f877214ee672093b52f07897ad90 |
| genomic_resource.yaml | 3.14 KB | b511f2daedbf62cd7518cad94741e9f2 |
| genomic_resource_original.yaml | 3.05 KB | aa52d42f56f190cf4b9c4faf1a88f46b |
| statistics/ |