TF_ChIP-seq_ENCSR000BTF
| Id: | TF_ChIP-seq/ENCSR000BTF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BTF [biosamplesummary="Homo sapiens HL-60" and target="REST"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN334XUB|/analyses/ENCAN334XUB/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_not_compliant: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF128UOK|/files/ENCFF128UOK/}, {ENCFF603MCO|/files/ENCFF603MCO/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.18 and a self consistency ratio of 4.32. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_not_compliant: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF975OED|/files/ENCFF975OED/}, {ENCFF939MDW|/files/ENCFF939MDW/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.18 and a self consistency ratio of 4.32. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Processed alignments file {ENCFF027GJA|/files/ENCFF027GJA/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19132039 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting REST-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
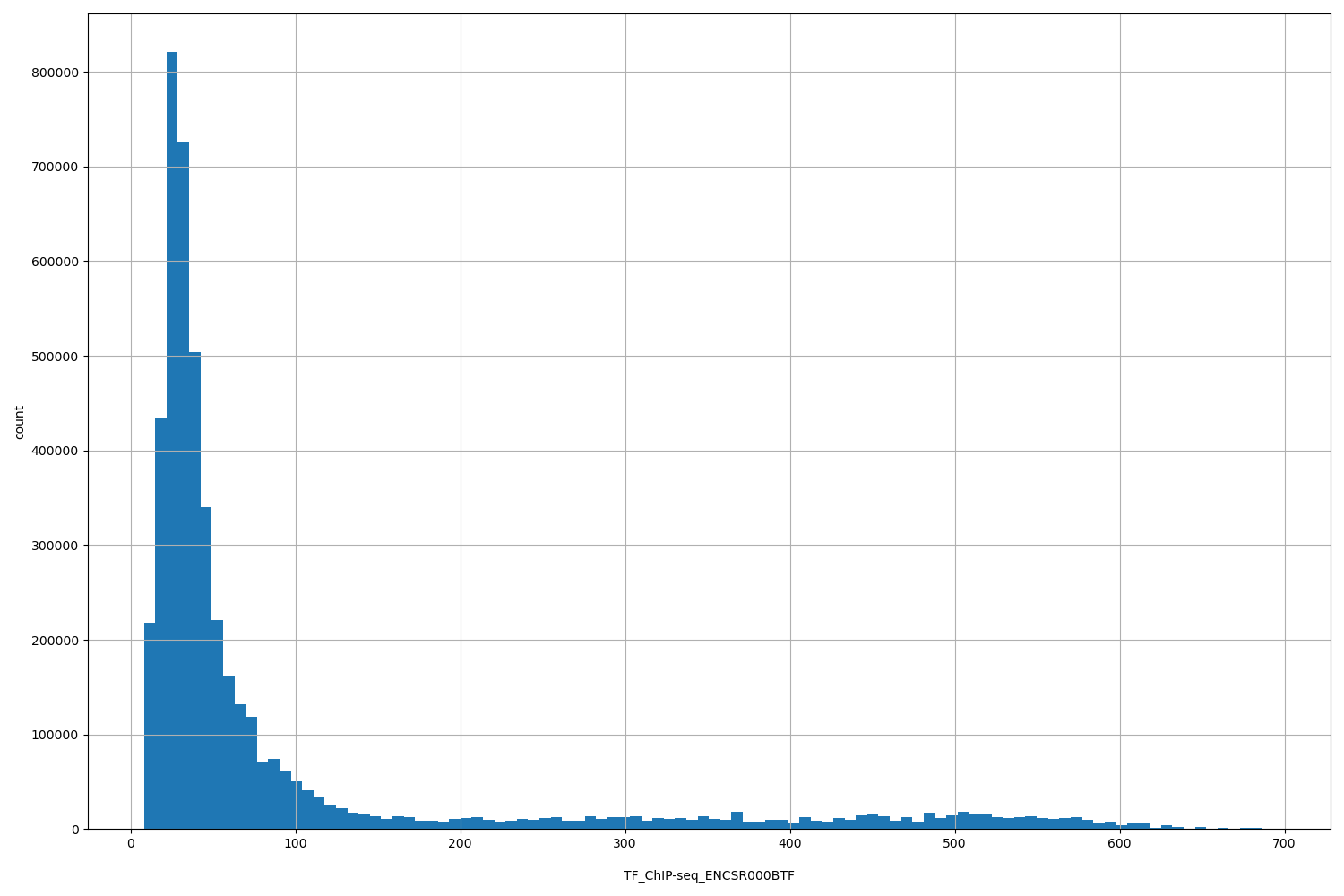
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BTF | float |
TF_ChIP-seq_ENCSR000BTF |
TF_ChIP-seq ENCSR000BTF [biosample_summary="Homo sapiens HL-60" and target="REST"]
|
 |
[8.11, 694] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF603MCO.bed.gz | 244.39 KB | bd65a9b594a044fa6e72d0baca077e59 |
| ENCFF603MCO.bed.gz.dvc | 100.0 B | 0f073f0d9920c62b8f5346263d9b41df |
| ENCFF603MCO.tabix.bed.gz | 177.12 KB | c892f57e372f8ed152a186ac288265ba |
| ENCFF603MCO.tabix.bed.gz.dvc | 106.0 B | d42462a04a14084ca235189226c3e8e3 |
| ENCFF603MCO.tabix.bed.gz.tbi | 102.49 KB | ef36f564f59850725e4e048ef0b95fd2 |
| ENCFF603MCO.tabix.bed.gz.tbi.dvc | 110.0 B | d92d03ec71d97c60eb07930ccfbe85dd |
| genomic_resource.yaml | 2.99 KB | 419a3fe3ee20487e17a2ffffa2834122 |
| genomic_resource_original.yaml | 2.89 KB | 3b24256ca2adec6f1a733ca9cd89597c |
| statistics/ |