TF_ChIP-seq_ENCSR000AMA
| Id: | TF_ChIP-seq/ENCSR000AMA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000AMA [biosamplesummary="Homo sapiens HepG2" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF129CNS|/files/ENCFF129CNS/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 1959659 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN293JLK|/analyses/ENCAN293JLK/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN293JLK|/analyses/ENCAN293JLK/} has in progress subobject document {1957f40f-d9d1-465b-be12-94371b07d22e|/documents/1957f40f-d9d1-465b-be12-94371b07d22e/} audit_warning: Processed alignments file {ENCFF520WEM|/files/ENCFF520WEM/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 10128882 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF205OKL|/files/ENCFF205OKL/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.33 and a self consistency ratio of 2.98. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF194VBQ|/files/ENCFF194VBQ/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.33 and a self consistency ratio of 2.98. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
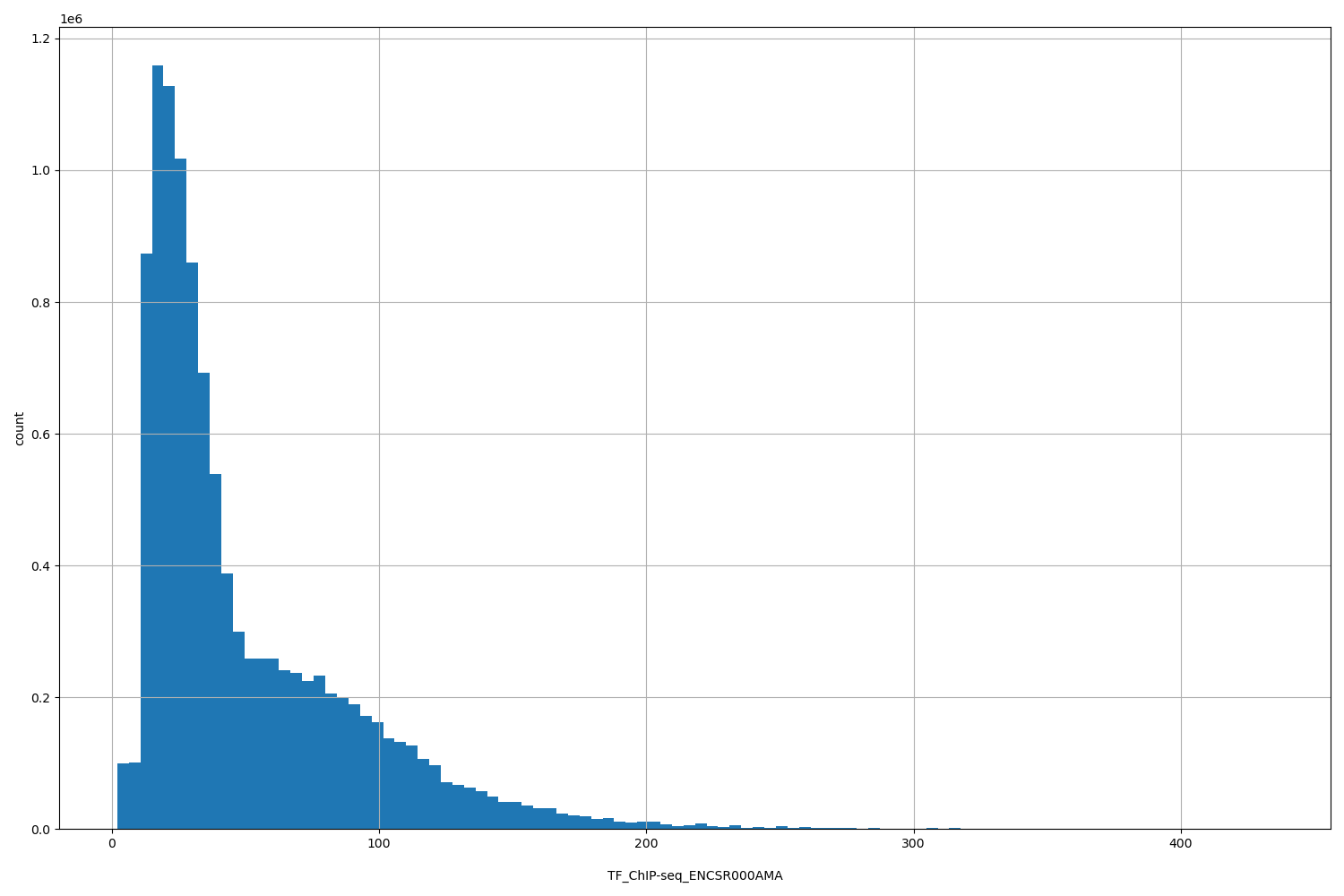
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000AMA | float |
TF_ChIP-seq_ENCSR000AMA |
TF_ChIP-seq ENCSR000AMA [biosample_summary="Homo sapiens HepG2" and target="CTCF"]
|
 |
[2, 434] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF205OKL.bed.gz | 713.3 KB | e8a6113a4bcdd9a73d5a9c0f6ada58d8 |
| ENCFF205OKL.bed.gz.dvc | 100.0 B | 15e23efa1eb53753bf8ca5db7d1f4ddc |
| ENCFF205OKL.tabix.bed.gz | 543.32 KB | 46182d3a99172ca3f4e3dac367f0998b |
| ENCFF205OKL.tabix.bed.gz.dvc | 106.0 B | ff499d3d08df33962cc6df5b57de57e6 |
| ENCFF205OKL.tabix.bed.gz.tbi | 288.36 KB | 1176ea1c988fbc2df19e66073625b8e5 |
| ENCFF205OKL.tabix.bed.gz.tbi.dvc | 110.0 B | 5bfece02a50cddce94a46dd379d6eb43 |
| genomic_resource.yaml | 3.75 KB | d4c6479a8a1f77f9a41d94844c4d8863 |
| genomic_resource_original.yaml | 3.65 KB | bd199a6e9e643b97fdbb3d7897531fdf |
| statistics/ |