Histone_ChIP-seq_ENCSR516KSY
| Id: | Histone_ChIP-seq/ENCSR516KSY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR516KSY [biosamplesummary="Homo sapiens H9 stably expressing HES5" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: stably expressing C-terminal eGFP-tagged HES5 output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN331XVJ|/analyses/ENCAN331XVJ/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN331XVJ|/analyses/ENCAN331XVJ/} has in progress subobject document {64024907-19d8-4c45-8bf9-869d4b5ab7db|/documents/64024907-19d8-4c45-8bf9-869d4b5ab7db/} audit_not_compliant: Processed unfiltered alignments file {ENCFF260JGU|/files/ENCFF260JGU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 25051464 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed unfiltered alignments file {ENCFF360AAU|/files/ENCFF360AAU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 41877987 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
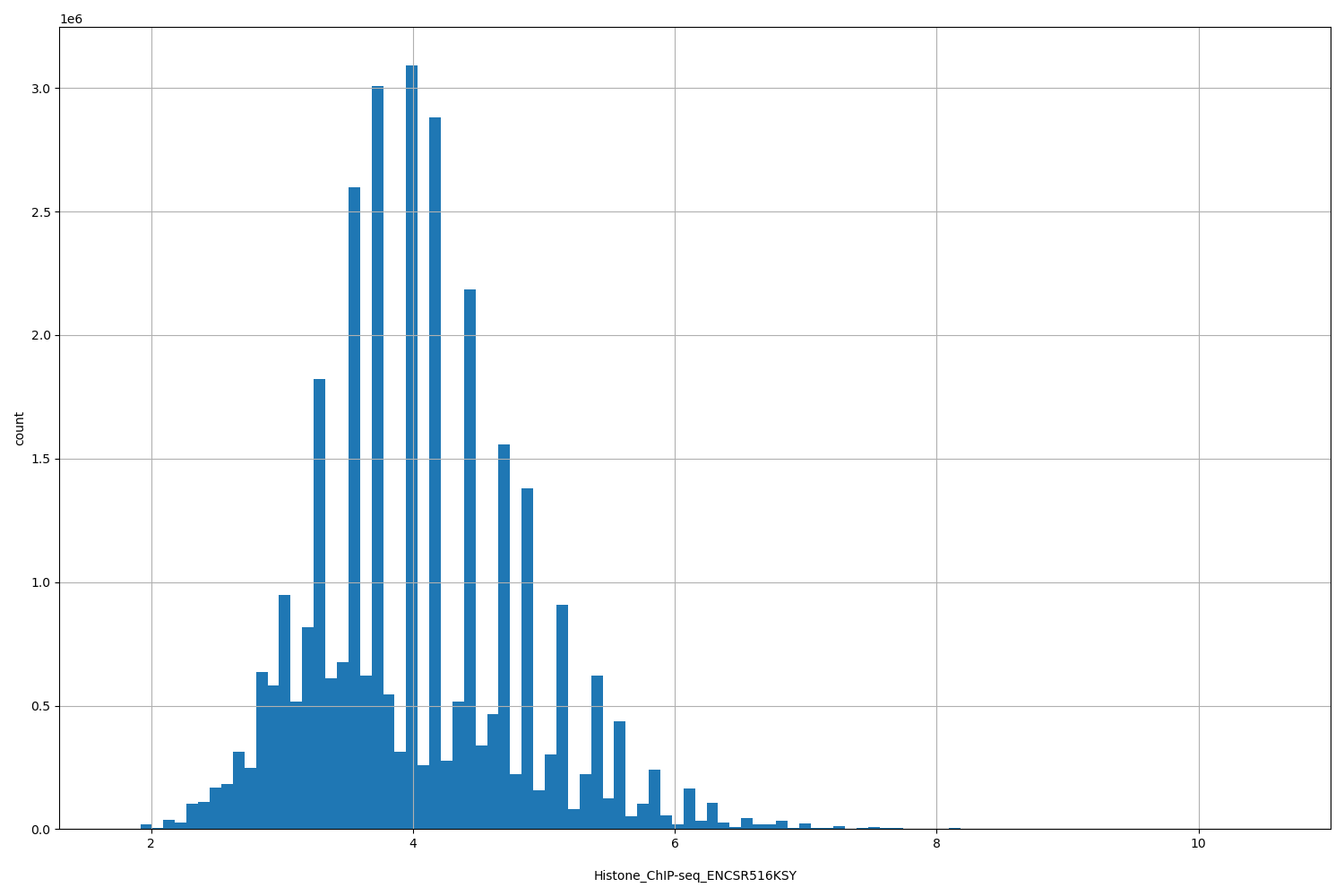
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR516KSY | float |
Histone_ChIP-seq_ENCSR516KSY |
Histone_ChIP-seq ENCSR516KSY [biosample_summary="Homo sapiens H9 stably expressing HES5" and target="H3K9me3"]
|
 |
[1.74, 10.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF724VHG.bed.gz | 905.17 KB | da754663dbb7d2273c596ad082b00593 |
| ENCFF724VHG.bed.gz.dvc | 100.0 B | e7bcacca6101aa1e8f382f523d469a59 |
| ENCFF724VHG.tabix.bed.gz | 476.78 KB | 5225038945e335801a8a6991759cec67 |
| ENCFF724VHG.tabix.bed.gz.dvc | 106.0 B | 98dfc347aa292a72064c4811a241d877 |
| ENCFF724VHG.tabix.bed.gz.tbi | 156.04 KB | e88560a7317efb3a117ee7db74e30996 |
| ENCFF724VHG.tabix.bed.gz.tbi.dvc | 110.0 B | 61fd237571b19e57ea65c85bc0c7b6c4 |
| genomic_resource.yaml | 2.7 KB | fa0c459a2405c4f341000033434cb366 |
| genomic_resource_original.yaml | 2.57 KB | 88443abf8fb1d4d9bd59c79cdbac82ca |
| statistics/ |