Histone_ChIP-seq_ENCSR442DGE
| Id: | Histone_ChIP-seq/ENCSR442DGE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR442DGE [biosamplesummary="Homo sapiens H1" and target="H3K79me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 3 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN458KFC|/analyses/ENCAN458KFC/} has in progress subobject document {8c7261b4-c230-4291-a833-77578ca69477|/documents/8c7261b4-c230-4291-a833-77578ca69477/} audit_internal_action: Released analysis {ENCAN458KFC|/analyses/ENCAN458KFC/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed alignments file {ENCFF341KSR|/files/ENCFF341KSR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 11460222 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF535GVC|/files/ENCFF535GVC/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16014785 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
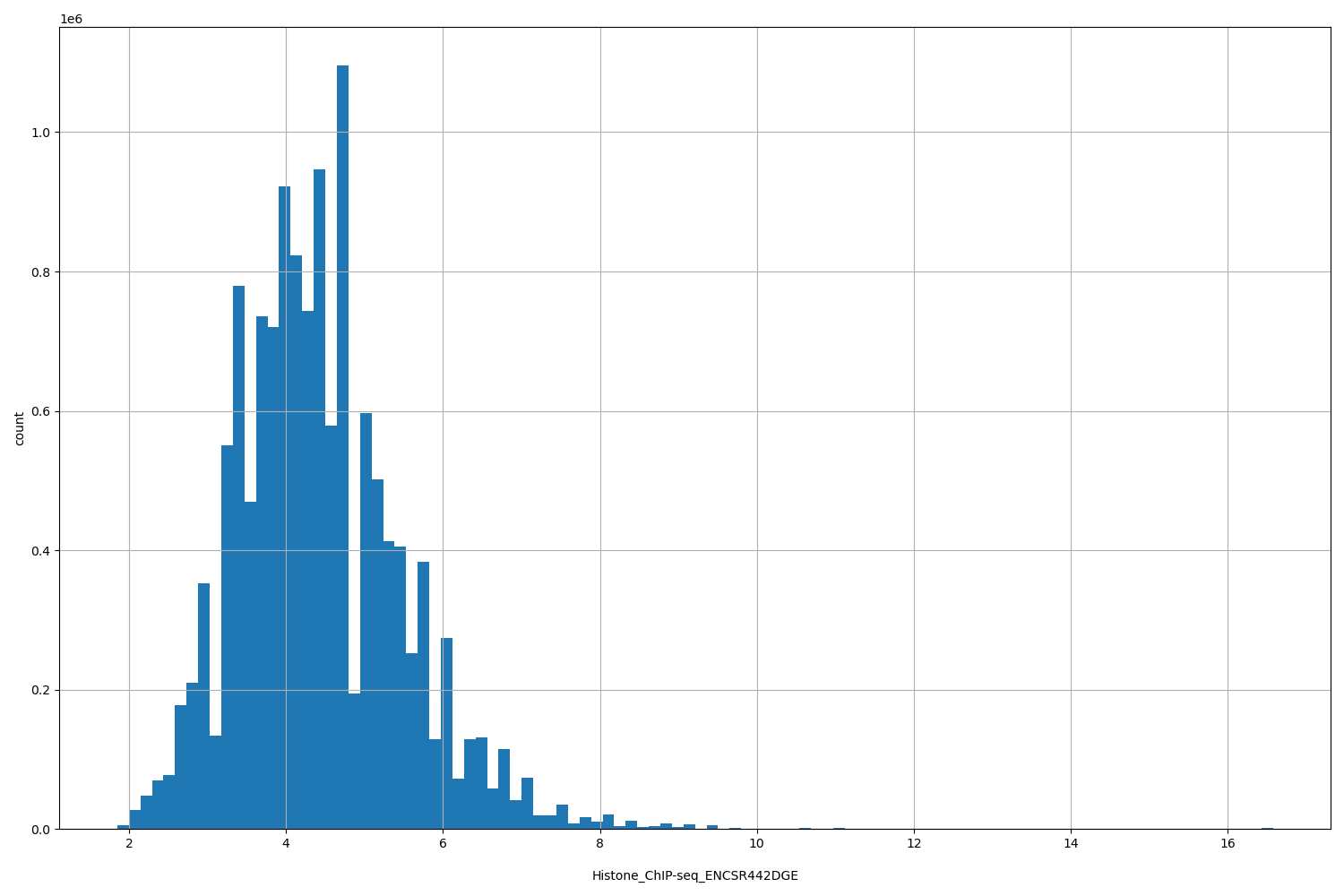
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR442DGE | float |
Histone_ChIP-seq_ENCSR442DGE |
Histone_ChIP-seq ENCSR442DGE [biosample_summary="Homo sapiens H1" and target="H3K79me1"]
|
 |
[1.85, 16.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF088PTH.bed.gz | 713.99 KB | 871a24d2369a317e1b1d31f7bae960fe |
| ENCFF088PTH.bed.gz.dvc | 100.0 B | 1496ed1379bf5988aa4d0f644f070b31 |
| ENCFF088PTH.tabix.bed.gz | 372.78 KB | 38d0cf751da03ab797a3febca8849416 |
| ENCFF088PTH.tabix.bed.gz.dvc | 106.0 B | 349e16ad272641e29bf48362c3eb38f2 |
| ENCFF088PTH.tabix.bed.gz.tbi | 130.59 KB | 0ae629319027669a0d86434aeafa3399 |
| ENCFF088PTH.tabix.bed.gz.tbi.dvc | 110.0 B | 16d7bbe860ce0c19d0355b5814f6a407 |
| genomic_resource.yaml | 2.71 KB | b3b41b8af6776bcd1c21f6111adfe5d8 |
| genomic_resource_original.yaml | 2.61 KB | 99ef3134cada317762f58bff3c3a7ecf |
| statistics/ |