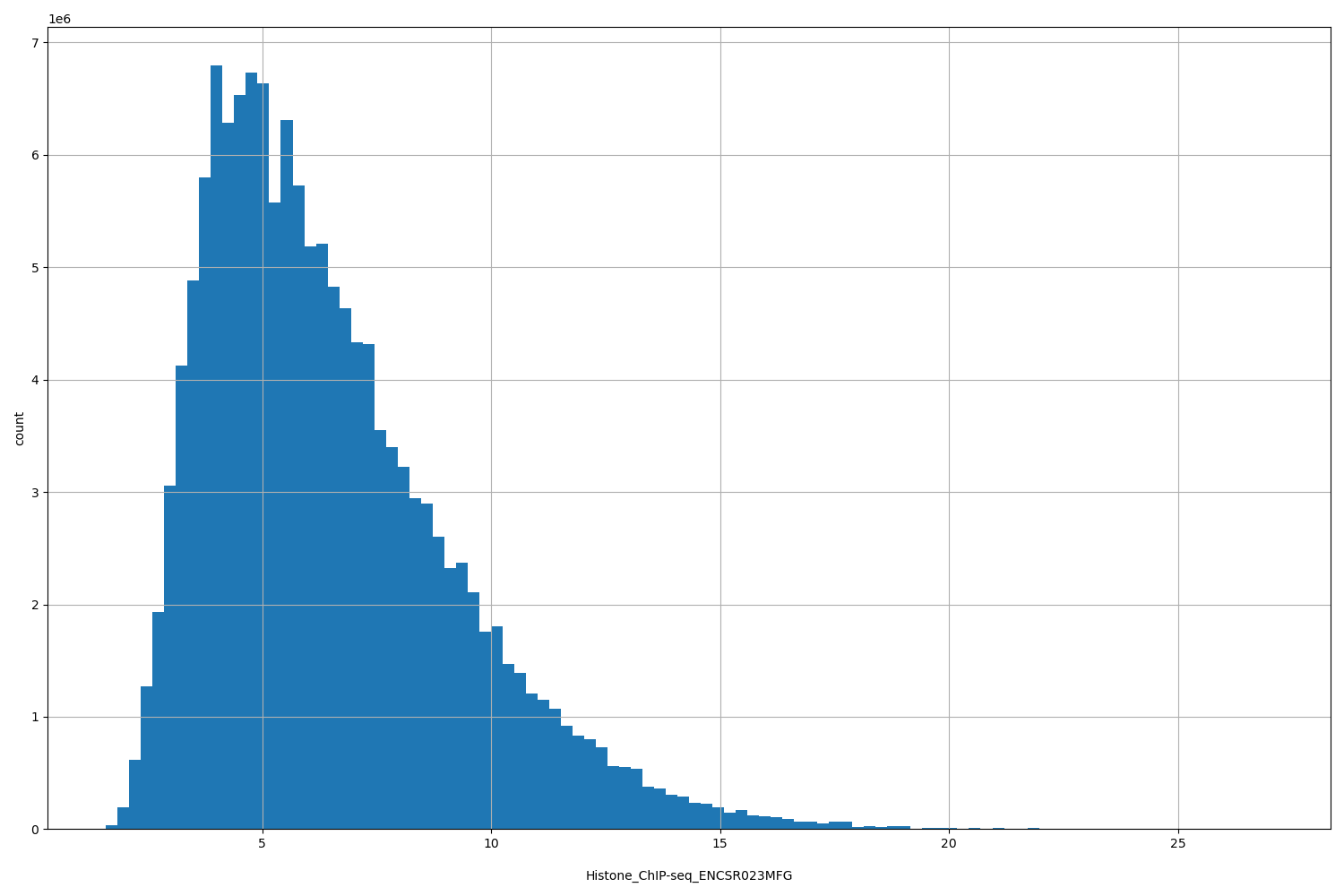
Histone_ChIP-seq_ENCSR023MFG
| Id: | Histone_ChIP-seq/ENCSR023MFG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR023MFG [biosamplesummary="Homo sapiens skeletal muscle satellite cell female adult originated from mesodermal cell" and target="H3K4me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 3 summary: female adult output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN989MDJ|/analyses/ENCAN989MDJ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed redacted alignments file {ENCFF647TSZ|/files/ENCFF647TSZ/} processed by ChIP-seq ENCODE3 hg19 pipeline has 17274025 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed redacted alignments file {ENCFF367QMK|/files/ENCFF367QMK/} processed by ChIP-seq ENCODE3 hg19 pipeline has 21770509 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR023MFG | float |
Histone_ChIP-seq_ENCSR023MFG |
Histone_ChIP-seq ENCSR023MFG [biosample_summary="Homo sapiens skeletal muscle satellite cell female adult originated from mesodermal cell" and target="H3K4me1"]
|
 |
[1.59, 27.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF428WDD.bed.gz | 4.02 MB | 5e63c3e81113eebbe5473ab128150bec |
| ENCFF428WDD.bed.gz.dvc | 101.0 B | ffca3f0f3dd72b2aec78f7e3f55c936a |
| ENCFF428WDD.tabix.bed.gz | 1.75 MB | 07d68b53db3c86fab1d3db8d2077c5cd |
| ENCFF428WDD.tabix.bed.gz.dvc | 107.0 B | 655379eb11672a983c47846593c0c065 |
| ENCFF428WDD.tabix.bed.gz.tbi | 386.93 KB | 41efe5a6f2a55ebd762694ff26aa4603 |
| ENCFF428WDD.tabix.bed.gz.tbi.dvc | 110.0 B | 386286e9614120a38a8f8b4679f1f74b |
| genomic_resource.yaml | 2.7 KB | a98f70ec3d050c3e1f41a082c9af06ab |
| genomic_resource_original.yaml | 2.53 KB | 3a8af48a36618a2734a4981c083f3652 |
| statistics/ |