Histone_ChIP-seq_ENCSR000FCP
| Id: | Histone_ChIP-seq/ENCSR000FCP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000FCP [biosamplesummary="Homo sapiens HCT116" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN829DUI|/analyses/ENCAN829DUI/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN829DUI|/analyses/ENCAN829DUI/} has in progress subobject document {74cb5647-23f1-4e2d-924b-0f9f92f1a038|/documents/74cb5647-23f1-4e2d-924b-0f9f92f1a038/} audit_warning: Processed unfiltered alignments file {ENCFF383RZK|/files/ENCFF383RZK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 37291408 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed unfiltered alignments file {ENCFF027DUB|/files/ENCFF027DUB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 36320111 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
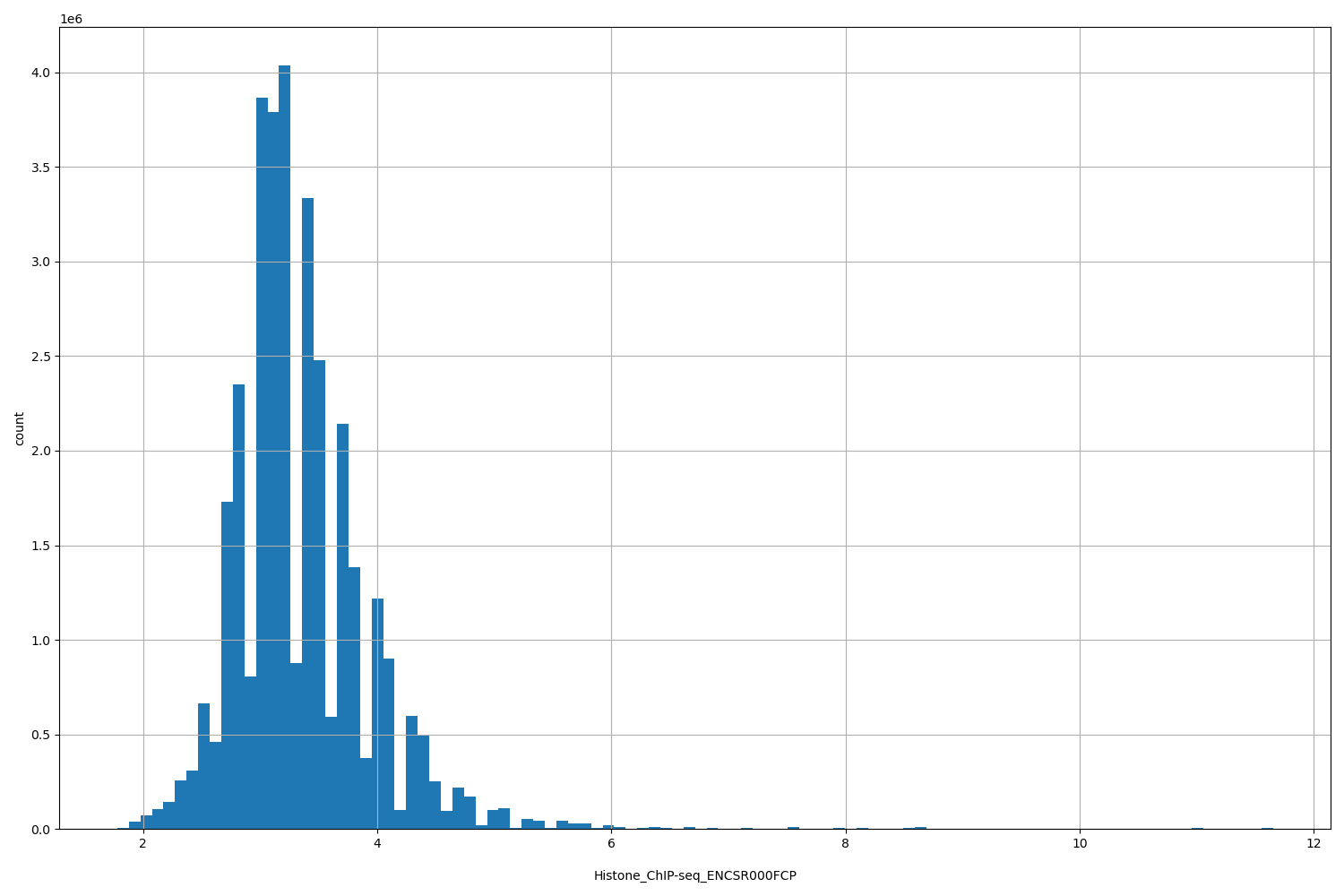
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000FCP | float |
Histone_ChIP-seq_ENCSR000FCP |
Histone_ChIP-seq ENCSR000FCP [biosample_summary="Homo sapiens HCT116" and target="H3K9me3"]
|
 |
[1.78, 11.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF158YTR.bed.gz | 1017.66 KB | 928edd5adf5ae87a0ebcad8c1652a7ef |
| ENCFF158YTR.bed.gz.dvc | 101.0 B | de75d254e4a123c8ef419a4d43dabb66 |
| ENCFF158YTR.tabix.bed.gz | 557.41 KB | 5ac5f9c5c3e646dbd777f0b934f8430e |
| ENCFF158YTR.tabix.bed.gz.dvc | 106.0 B | eb532fea28ed9dfe4051baac7df382cb |
| ENCFF158YTR.tabix.bed.gz.tbi | 173.91 KB | ae319ed107e603b877ac0517f0afbb3e |
| ENCFF158YTR.tabix.bed.gz.tbi.dvc | 110.0 B | c7b2d8aae73c54dd7d6a91c172511008 |
| genomic_resource.yaml | 2.66 KB | a66220b98355d90524e2362f2241ba3d |
| genomic_resource_original.yaml | 2.55 KB | 60fa3a07bd3d0a63d1b0dd47921b93cf |
| statistics/ |