DNase-seq_ENCSR000EQC
| Id: | DNase-seq/ENCSR000EQC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EQC [biosample_summary="Homo sapiens T-helper 1 cell"] |
| Description: |
status: released biological_replicates: Rep 2 summary: output_type: peaks audit_internal_action: Archived analysis {ENCAN305CDI|/analyses/ENCAN305CDI/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_not_compliant: Replicate concordance in DNase-seq experiments is measured by calculating the Pearson correlation between signal quantification of the replicates. ENCODE processed signal files {ENCFF479VHW|/files/ENCFF479VHW/}, {ENCFF148BTW|/files/ENCFF148BTW/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL001DNS|/pipelines/ENCPL001DNS/}) for GRCh38 assembly have a Pearson correlation of 0.53. According to ENCODE standards, in an isogenic assay a Pearson correlation value > 0.9 is recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/} ) |
| Labels: |
|
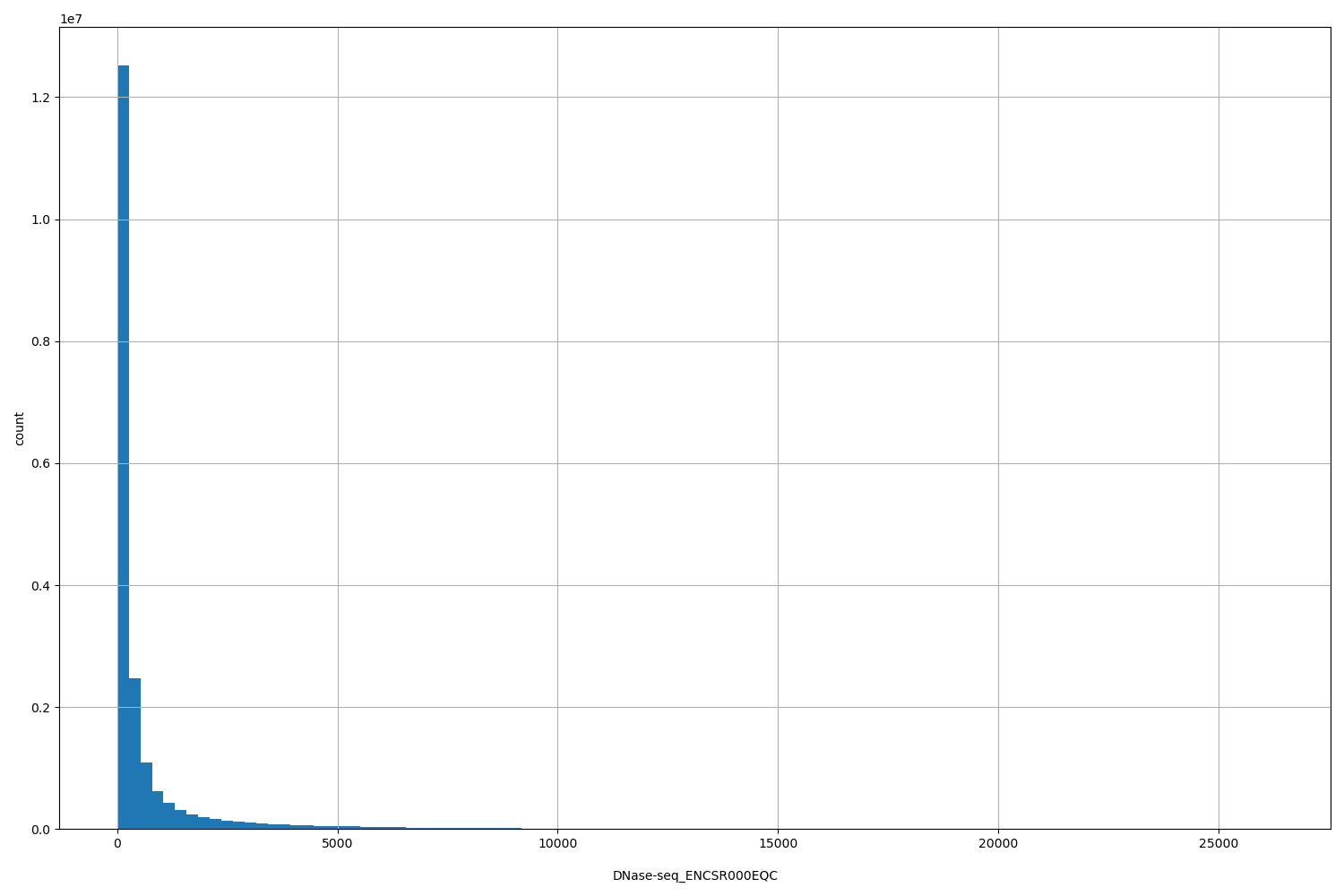
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EQC | float |
DNase-seq_ENCSR000EQC |
DNase-seq ENCSR000EQC [biosample_summary="Homo sapiens T-helper 1 cell"]
|
 |
[3, 2.62e+04] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF920ELS.bed.gz | 1.27 MB | d79937a88e830880f8075dc497eb085c |
| ENCFF920ELS.bed.gz.dvc | 101.0 B | 75a02cecbf72bc89aaf4bab9b3b34871 |
| ENCFF920ELS.tabix.bed.gz | 1.1 MB | 1f946e01432d289cd60328adee1e676f |
| ENCFF920ELS.tabix.bed.gz.dvc | 107.0 B | 27a1d3a5ff16030d9cad2ddea787a795 |
| ENCFF920ELS.tabix.bed.gz.tbi | 465.52 KB | 49af1c946ba5fa47d7dbad97e0bc6860 |
| ENCFF920ELS.tabix.bed.gz.tbi.dvc | 110.0 B | 40c17e470e2777064fecf1d79d4a8904 |
| genomic_resource.yaml | 1.91 KB | e6c1d32640425aa0dc26efbcf3e48b7b |
| genomic_resource_original.yaml | 1.83 KB | c049aade50fe929640ed17087f01b881 |
| statistics/ |