ATAC-seq_ENCSR846VPV
| Id: | ATAC-seq/ENCSR846VPV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR846VPV [biosample_summary="Homo sapiens heart left ventricle tissue female adult (66 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (66 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN380SYM|/analyses/ENCAN380SYM/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_internal_action: Released analysis {ENCAN380SYM|/analyses/ENCAN380SYM/} has in progress subobject document {1ffc1b32-5e74-41f8-9011-363a45567cd6|/documents/1ffc1b32-5e74-41f8-9011-363a45567cd6/} audit_not_compliant: Alignment file {ENCFF456PVX|/files/ENCFF456PVX/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has 17070881 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF456PVX|/files/ENCFF456PVX/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.881. |
| Labels: |
|
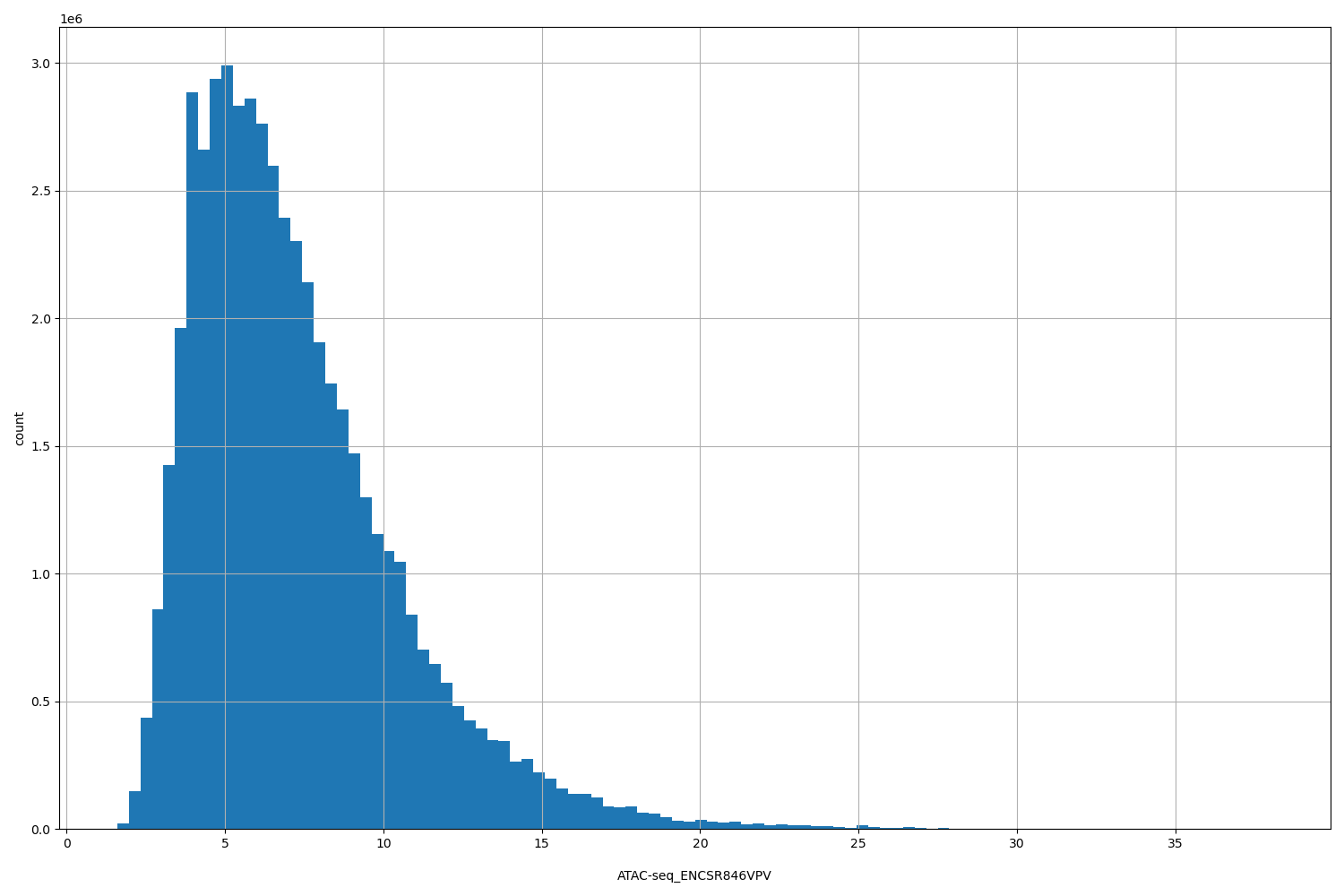
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR846VPV | float |
ATAC-seq_ENCSR846VPV |
ATAC-seq ENCSR846VPV [biosample_summary="Homo sapiens heart left ventricle tissue female adult (66 years)"]
|
 |
[1.6, 38.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF085ASE.bed.gz | 3.77 MB | 44fec0fcb22c81941c2dcda4a6ffd139 |
| ENCFF085ASE.bed.gz.dvc | 101.0 B | 0b48ec243a6bde0be8420ad0a145f43a |
| ENCFF085ASE.tabix.bed.gz | 1.14 MB | 528ffbd4838ff23df8ede9b82ffd97b2 |
| ENCFF085ASE.tabix.bed.gz.dvc | 107.0 B | 33be1eba61ac4dfd15a84ed8d8693202 |
| ENCFF085ASE.tabix.bed.gz.tbi | 431.93 KB | 0ecac038efdd81352c75b8b40a485904 |
| ENCFF085ASE.tabix.bed.gz.tbi.dvc | 110.0 B | e2862ae6f209bf86257f2cfa2d51e69e |
| genomic_resource.yaml | 2.54 KB | 1854b78b533c239bf15c997840182acf |
| genomic_resource_original.yaml | 2.43 KB | e38429c3ff2eb461ed3f615d315526a8 |
| statistics/ |