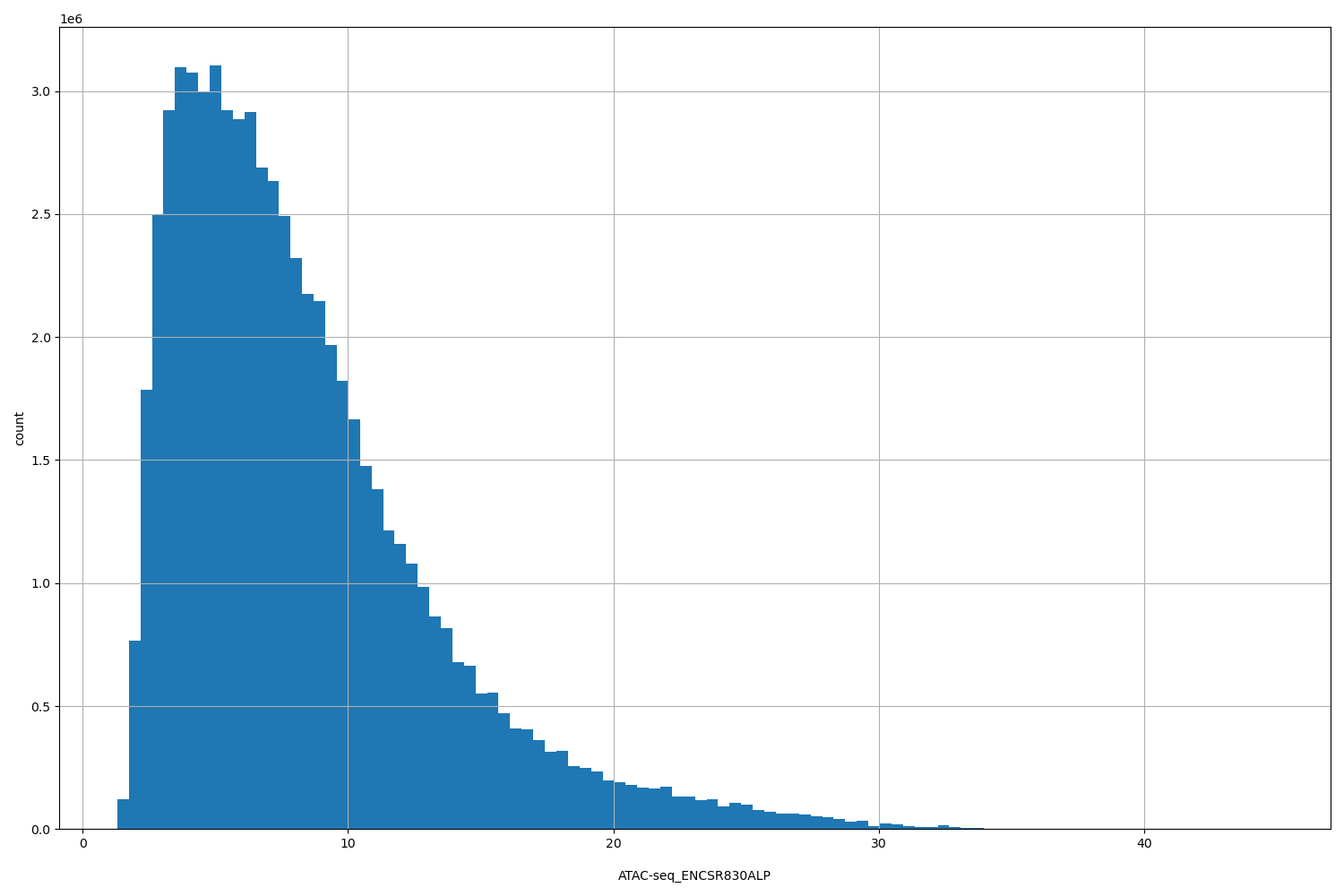
ATAC-seq_ENCSR830ALP
| Id: | ATAC-seq/ENCSR830ALP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR830ALP [biosample_summary="Homo sapiens activated T-cell male adult (43 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (43 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN041QSD|/analyses/ENCAN041QSD/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF466GWI|/files/ENCFF466GWI/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with NRF value of 0.891593. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF910SFN|/files/ENCFF910SFN/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with NRF value of 0.893802. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR830ALP | float |
ATAC-seq_ENCSR830ALP |
ATAC-seq ENCSR830ALP [biosample_summary="Homo sapiens activated T-cell male adult (43 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours"]
|
 |
[1.31, 44.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF350JTS.bed.gz | 4.56 MB | 245bbbafa6541cca487cca6d58ea0f8b |
| ENCFF350JTS.bed.gz.dvc | 101.0 B | c8552b6cf89e050daef00cac2c065dc1 |
| ENCFF350JTS.tabix.bed.gz | 1.26 MB | 674423ff3a2ff51e40db43e24c4b39f5 |
| ENCFF350JTS.tabix.bed.gz.dvc | 107.0 B | 5db6543457e7d4fee3fd56805e7e8c24 |
| ENCFF350JTS.tabix.bed.gz.tbi | 424.12 KB | 98c7cff58b6862200f94225068fcadc3 |
| ENCFF350JTS.tabix.bed.gz.tbi.dvc | 110.0 B | 184d561765be429002dcf96f6a6537db |
| genomic_resource.yaml | 2.94 KB | 459ca8f7a016a1fa80bca522481fbaef |
| genomic_resource_original.yaml | 2.76 KB | d04497974e6ec315795ceda36b6a1828 |
| statistics/ |