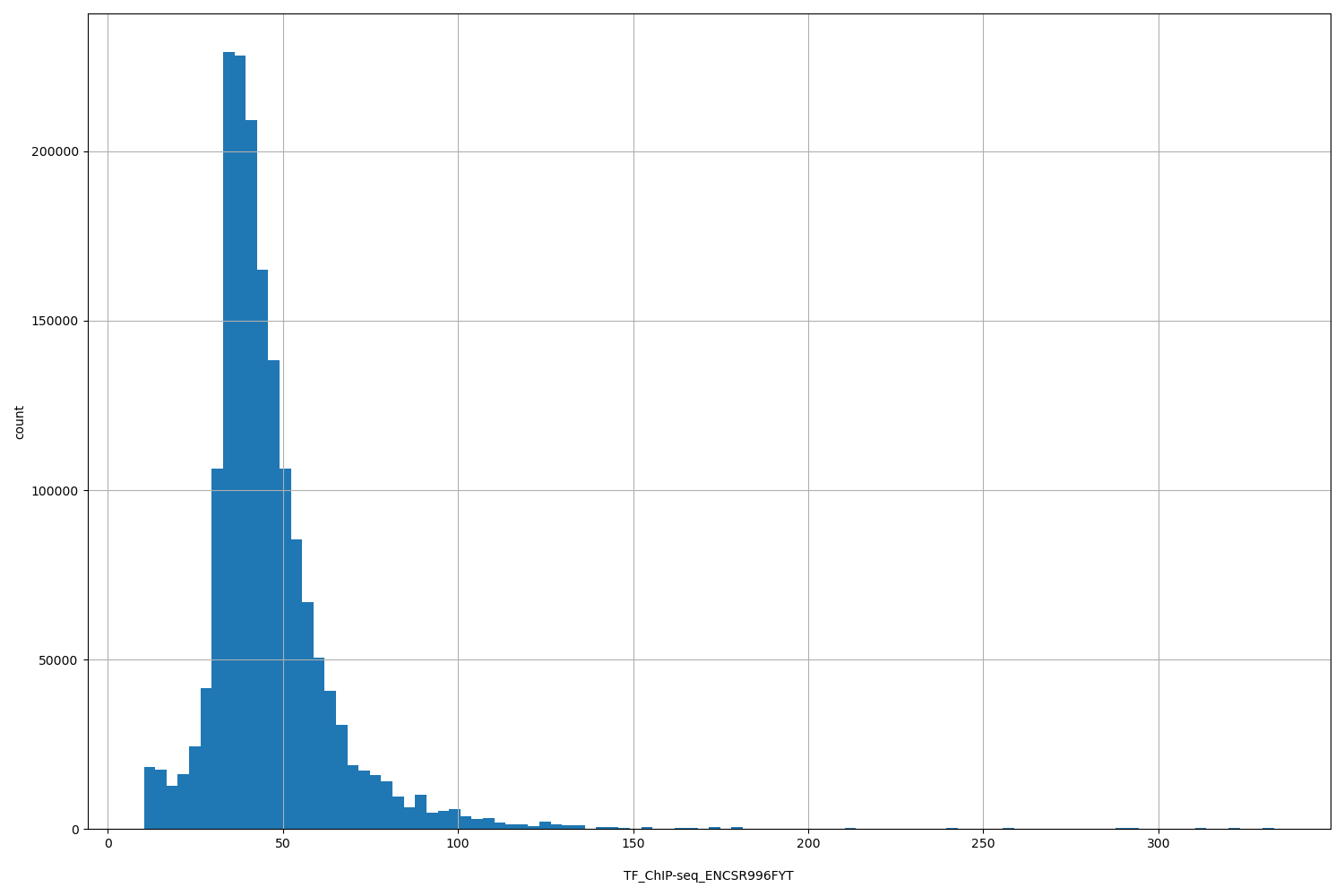
TF_ChIP-seq_ENCSR996FYT
| Id: | TF_ChIP-seq/ENCSR996FYT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR996FYT [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF202" and target="ZNF202"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF202 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN421MNO|/analyses/ENCAN421MNO/} has in progress subobject document {e29bf756-ca2a-4f58-9d69-0b65f2c2bd2d|/documents/e29bf756-ca2a-4f58-9d69-0b65f2c2bd2d/} audit_internal_action: Released analysis {ENCAN421MNO|/analyses/ENCAN421MNO/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF284XHX|/files/ENCFF284XHX/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.25 and a self consistency ratio of 2.94. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF574FZA|/files/ENCFF574FZA/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.25 and a self consistency ratio of 2.94. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR996FYT | float |
TF_ChIP-seq_ENCSR996FYT |
TF_ChIP-seq ENCSR996FYT [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF202" and target="ZNF202"]
|
 |
[10.4, 333] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF284XHX.bed.gz | 106.9 KB | 732e176360d4e89ba4deb5c7c8e695d3 |
| ENCFF284XHX.bed.gz.dvc | 100.0 B | 9f01e1a68b12bab940e3bdd20be6d4d8 |
| ENCFF284XHX.tabix.bed.gz | 70.04 KB | 6ce355b0f2632d0b5e55408ed4a4cf85 |
| ENCFF284XHX.tabix.bed.gz.dvc | 105.0 B | d0de6dd08120cf04fae12019110c7990 |
| ENCFF284XHX.tabix.bed.gz.tbi | 52.86 KB | 9faa7f088a1f64a2d5e50d41750a31c8 |
| ENCFF284XHX.tabix.bed.gz.tbi.dvc | 109.0 B | 37a9e1fcf9352a546e1826083399554e |
| genomic_resource.yaml | 3.07 KB | 78ae9fe10428a2d8271204a0b8704cd0 |
| genomic_resource_original.yaml | 2.88 KB | ebf2df69e602db6b256af3bdb60d6c1d |
| statistics/ |