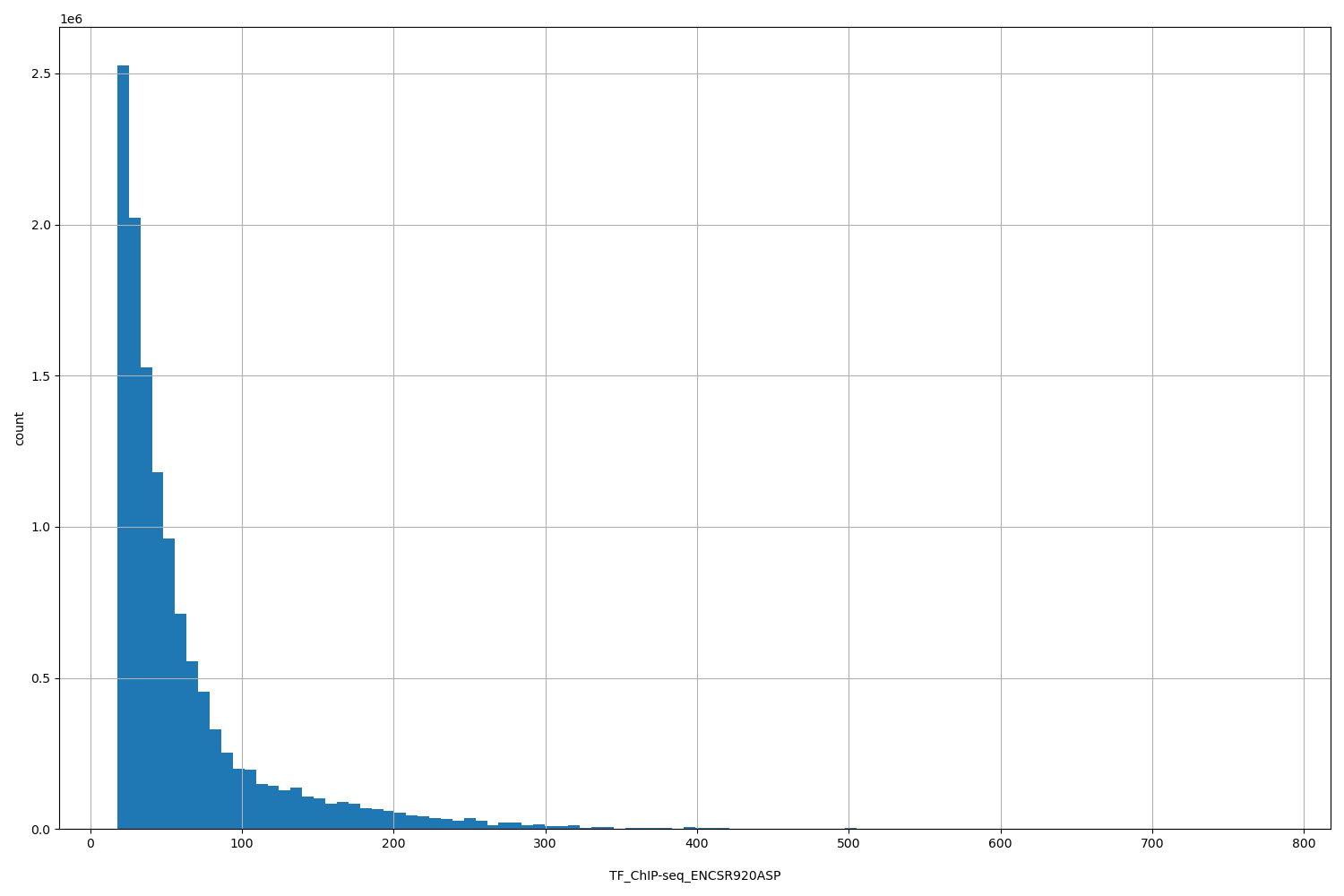
TF_ChIP-seq_ENCSR920ASP
| Id: | TF_ChIP-seq/ENCSR920ASP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR920ASP [biosamplesummary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZFX" and target="ZFX"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZFX output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN619YUE|/analyses/ENCAN619YUE/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF500GNW|/files/ENCFF500GNW/} processed by ChIP-seq ENCODE3 hg19 pipeline has 18027445 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZFX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF804GZU|/files/ENCFF804GZU/} processed by ChIP-seq ENCODE3 hg19 pipeline has 18804717 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZFX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR920ASP | float |
TF_ChIP-seq_ENCSR920ASP |
TF_ChIP-seq ENCSR920ASP [biosample_summary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZFX" and target="ZFX"]
|
 |
[17.9, 779] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF433RKB.bed.gz | 412.44 KB | f5ce515799a684a69f3d908d6f5358e2 |
| ENCFF433RKB.bed.gz.dvc | 100.0 B | 0b3707f1ba89532557ba23bf14152400 |
| ENCFF433RKB.tabix.bed.gz | 373.28 KB | 6cb17902980b6f165b409bda2a109294 |
| ENCFF433RKB.tabix.bed.gz.dvc | 106.0 B | 39c5ccfe2c89f8e2bf3ffe9723900224 |
| ENCFF433RKB.tabix.bed.gz.tbi | 127.78 KB | 30ea8ef1f984289c391be1e644a07de8 |
| ENCFF433RKB.tabix.bed.gz.tbi.dvc | 110.0 B | 2ab94845eb6afac5ba702aa23e0e13e1 |
| genomic_resource.yaml | 2.63 KB | 35edadafd7ae30216dd447d2ab18fd5f |
| genomic_resource_original.yaml | 2.48 KB | d3144e3a679c27416666da78b94e8b32 |
| statistics/ |