TF_ChIP-seq_ENCSR913ODG
| Id: | TF_ChIP-seq/ENCSR913ODG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR913ODG [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens MTF2" and target="MTF2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens MTF2 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN486ICU|/analyses/ENCAN486ICU/} has in progress subobject document {f01d40fd-c1f3-4ed7-b9a8-911f4a328ad4|/documents/f01d40fd-c1f3-4ed7-b9a8-911f4a328ad4/} audit_internal_action: Released analysis {ENCAN486ICU|/analyses/ENCAN486ICU/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF620YHG|/files/ENCFF620YHG/} processed by ChIP-seq ENCODE4 v1.5.0 GRCh38 pipeline have a rescue ratio of 1.47 and a self consistency ratio of 3.55. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF916FZN|/files/ENCFF916FZN/} processed by ChIP-seq ENCODE4 v1.5.0 GRCh38 pipeline have a rescue ratio of 1.47 and a self consistency ratio of 3.55. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
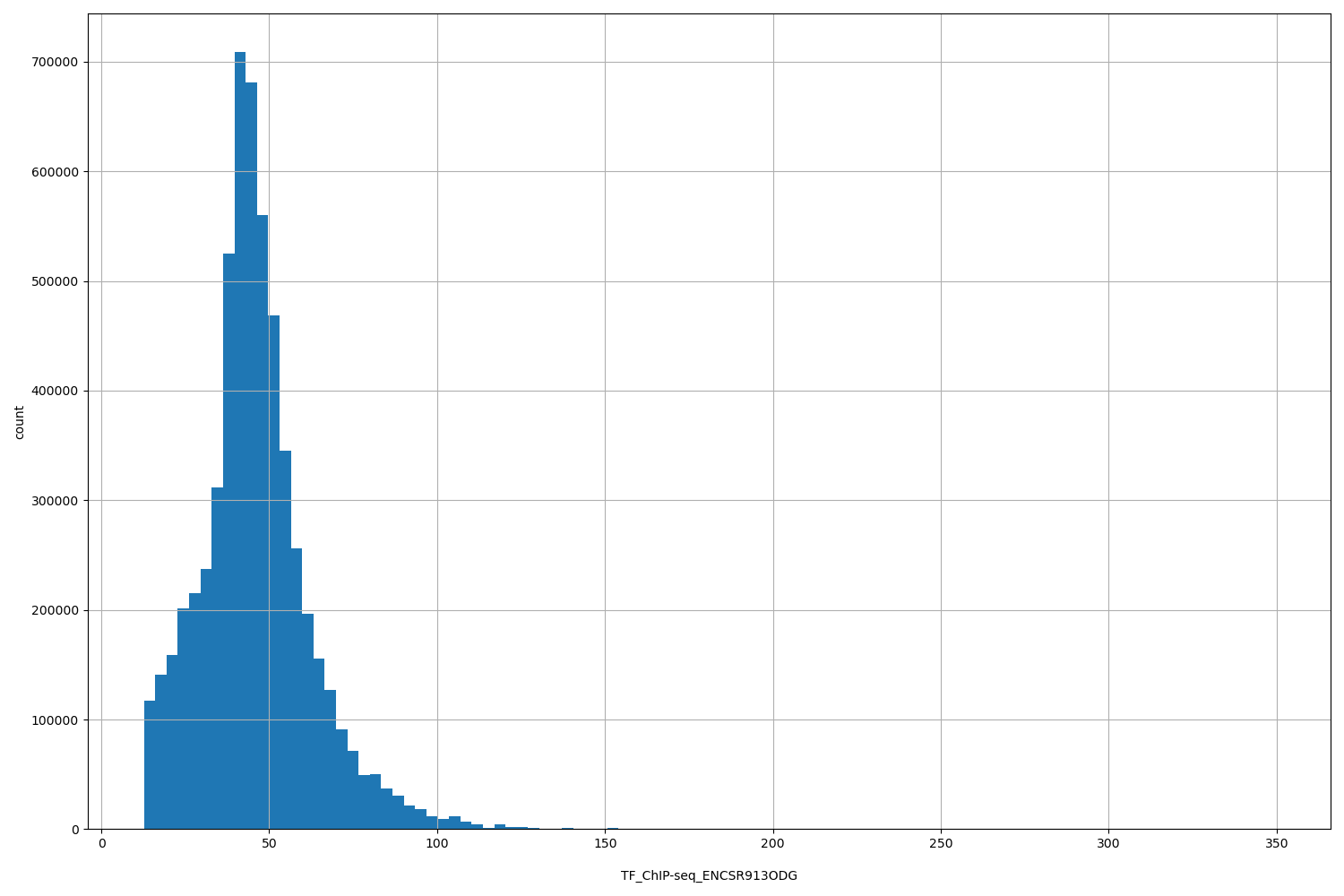
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR913ODG | float |
TF_ChIP-seq_ENCSR913ODG |
TF_ChIP-seq ENCSR913ODG [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens MTF2" and target="MTF2"]
|
 |
[12.7, 349] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF620YHG.bed.gz | 194.4 KB | c33704e20c1cfb3a6780d76a1f864604 |
| ENCFF620YHG.bed.gz.dvc | 100.0 B | b468d348614d3589895e65883cb9583c |
| ENCFF620YHG.tabix.bed.gz | 153.44 KB | caa09356a74da6fbb6f276e3289965d9 |
| ENCFF620YHG.tabix.bed.gz.dvc | 106.0 B | 0e25bc3766b63ae2a4ddf95ce0593e3b |
| ENCFF620YHG.tabix.bed.gz.tbi | 69.22 KB | fa05f472c8bd76eea8f7ce4c8645c855 |
| ENCFF620YHG.tabix.bed.gz.tbi.dvc | 109.0 B | 9520ec16acaa758afdede4efa8e8661c |
| genomic_resource.yaml | 2.94 KB | 5a689a8ba87ef799f8c9b41cf38e4c78 |
| genomic_resource_original.yaml | 2.78 KB | 152f721c9efb41013e6cacde19903ea4 |
| statistics/ |