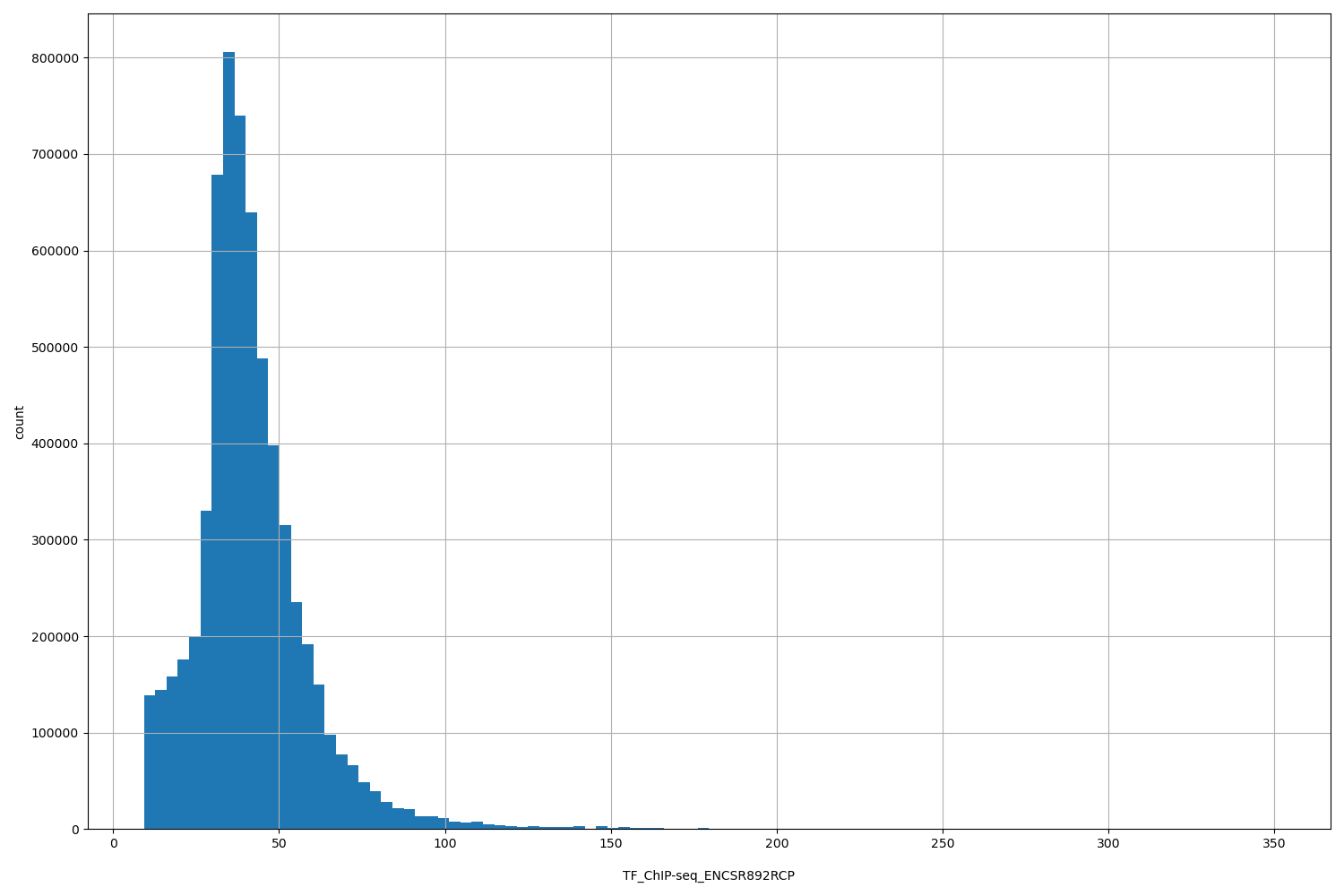
TF_ChIP-seq_ENCSR892RCP
| Id: | TF_ChIP-seq/ENCSR892RCP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR892RCP [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens CBX5" and target="CBX5"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens CBX5 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN364PCK|/analyses/ENCAN364PCK/} has in progress subobject document {d7a6af98-3102-40e3-b77a-a7ce4e1c8082|/documents/d7a6af98-3102-40e3-b77a-a7ce4e1c8082/} audit_internal_action: Released analysis {ENCAN364PCK|/analyses/ENCAN364PCK/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF486LNN|/files/ENCFF486LNN/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.50 and a self consistency ratio of 6.48. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF251YQZ|/files/ENCFF251YQZ/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.50 and a self consistency ratio of 6.48. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR892RCP | float |
TF_ChIP-seq_ENCSR892RCP |
TF_ChIP-seq ENCSR892RCP [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens CBX5" and target="CBX5"]
|
 |
[9.29, 350] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF486LNN.bed.gz | 298.53 KB | d18ff13cde5d77a76fff10b1cc0d47cb |
| ENCFF486LNN.bed.gz.dvc | 100.0 B | 38efd76b362c3ac0bbe1216015e9d50f |
| ENCFF486LNN.tabix.bed.gz | 235.12 KB | 37fce57959e53183a026adeba54e2391 |
| ENCFF486LNN.tabix.bed.gz.dvc | 106.0 B | 002a522aa2ae9707b6fc905bc0c04960 |
| ENCFF486LNN.tabix.bed.gz.tbi | 114.75 KB | e468c1a84b818dcc67d1df406fea4d3c |
| ENCFF486LNN.tabix.bed.gz.tbi.dvc | 110.0 B | d2325fe64582929e7321cec959f294e8 |
| genomic_resource.yaml | 2.94 KB | 49086cbe9acbe12c63afcf4e52fb8aff |
| genomic_resource_original.yaml | 2.78 KB | fb1a565dca0bb0600e445a906490b662 |
| statistics/ |