TF_ChIP-seq_ENCSR803PAI
| Id: | TF_ChIP-seq/ENCSR803PAI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR803PAI [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ATF4" and target="ATF4"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ATF4 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN909DIN|/analyses/ENCAN909DIN/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF056GCS|/files/ENCFF056GCS/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline has 18801116 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ATF4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF640PRA|/files/ENCFF640PRA/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline has 18690061 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ATF4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
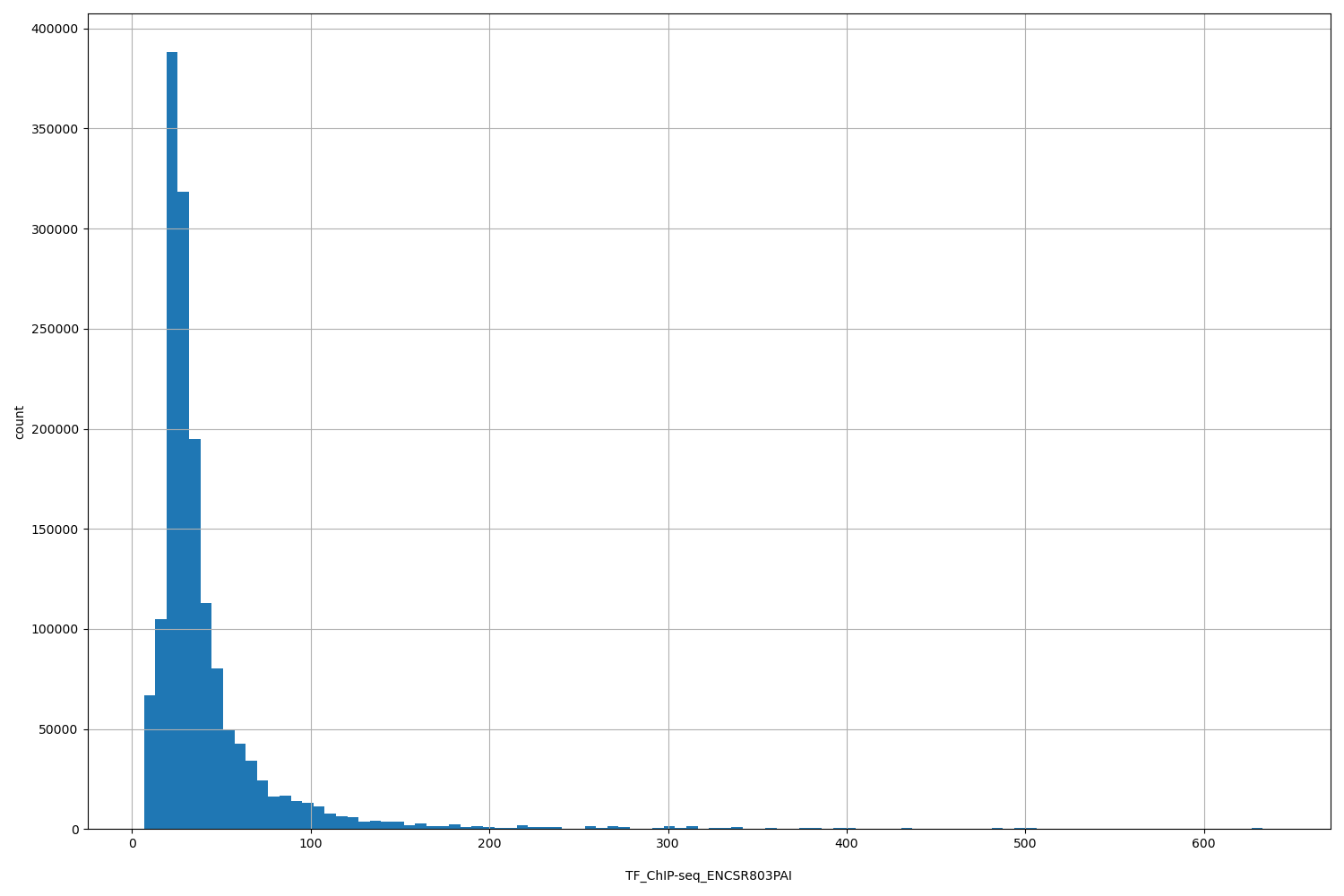
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR803PAI | float |
TF_ChIP-seq_ENCSR803PAI |
TF_ChIP-seq ENCSR803PAI [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ATF4" and target="ATF4"]
|
 |
[6.62, 639] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF885VKD.bed.gz | 86.45 KB | 07bf7d26c06d0cd9fdf031a6f3ab874c |
| ENCFF885VKD.bed.gz.dvc | 99.0 B | 039ce3388915d24592b3c4ce8a72840c |
| ENCFF885VKD.tabix.bed.gz | 60.75 KB | fa5d45d89b4409dec3f2f6163a68c346 |
| ENCFF885VKD.tabix.bed.gz.dvc | 105.0 B | 63788a16f0d61b77350fb91ca86bb7b3 |
| ENCFF885VKD.tabix.bed.gz.tbi | 44.18 KB | de02a8493f2da870973d796b29b362f6 |
| ENCFF885VKD.tabix.bed.gz.tbi.dvc | 109.0 B | 479bde9c5125dbf0a322c10ad10f92b4 |
| genomic_resource.yaml | 2.66 KB | 7d9b59dbfbff473e5ab853378502ef56 |
| genomic_resource_original.yaml | 2.5 KB | 42d6dcfe879a5c4bca93e0cd877920de |
| statistics/ |