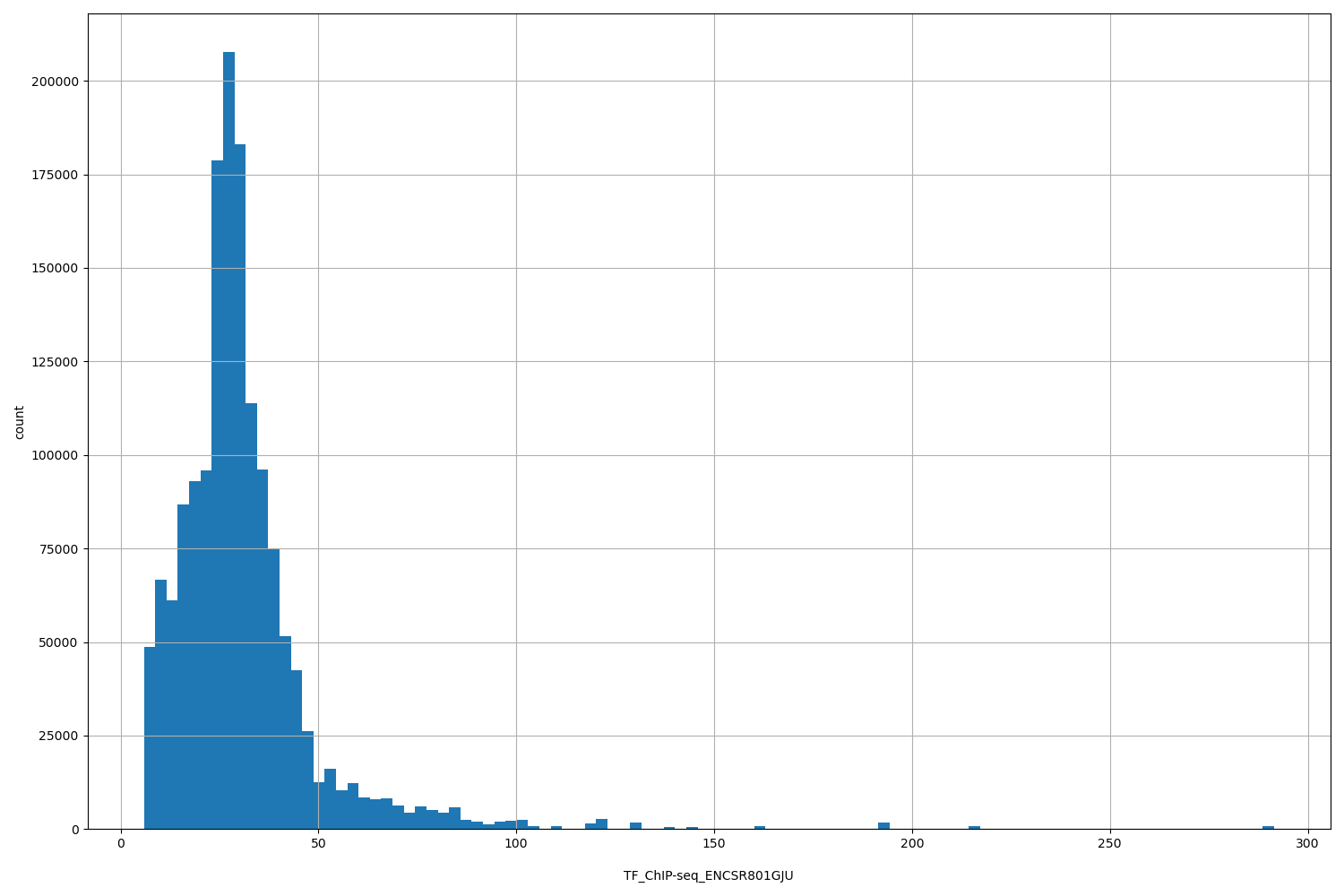
TF_ChIP-seq_ENCSR801GJU
| Id: | TF_ChIP-seq/ENCSR801GJU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR801GJU [biosamplesummary="Homo sapiens HepG2" and target="CBX2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF501QII|/files/ENCFF501QII/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN853SCC|/analyses/ENCAN853SCC/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF778SLS|/files/ENCFF778SLS/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 16973577 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CBX2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF978GYX|/files/ENCFF978GYX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 15209268 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CBX2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF501QII|/files/ENCFF501QII/}, {ENCFF027PQP|/files/ENCFF027PQP/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.00. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF111JJZ|/files/ENCFF111JJZ/}, {ENCFF027KUC|/files/ENCFF027KUC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.00. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR801GJU | float |
TF_ChIP-seq_ENCSR801GJU |
TF_ChIP-seq ENCSR801GJU [biosample_summary="Homo sapiens HepG2" and target="CBX2"]
|
 |
[5.9, 291] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF501QII.bed.gz | 54.41 KB | e59b618aa7d4737b773b09828807101a |
| ENCFF501QII.bed.gz.dvc | 99.0 B | ae0dbf67d9434bab2a6f5dea1ad66368 |
| ENCFF501QII.tabix.bed.gz | 44.02 KB | 46f8dd6e8e2dbac357b1ddf029e11c04 |
| ENCFF501QII.tabix.bed.gz.dvc | 105.0 B | 076eb3a0879c3d63614e2974752ca0cf |
| ENCFF501QII.tabix.bed.gz.tbi | 23.1 KB | 243fe00ea9012359788a6bfab5d885cd |
| ENCFF501QII.tabix.bed.gz.tbi.dvc | 109.0 B | 96c34bdec6d603fb79959f3cd5a0674d |
| genomic_resource.yaml | 3.65 KB | e9600e08d200ee77a1f79d699db6b215 |
| genomic_resource_original.yaml | 3.56 KB | 9b33c173a04d41d1f58287a29ceeb379 |
| statistics/ |