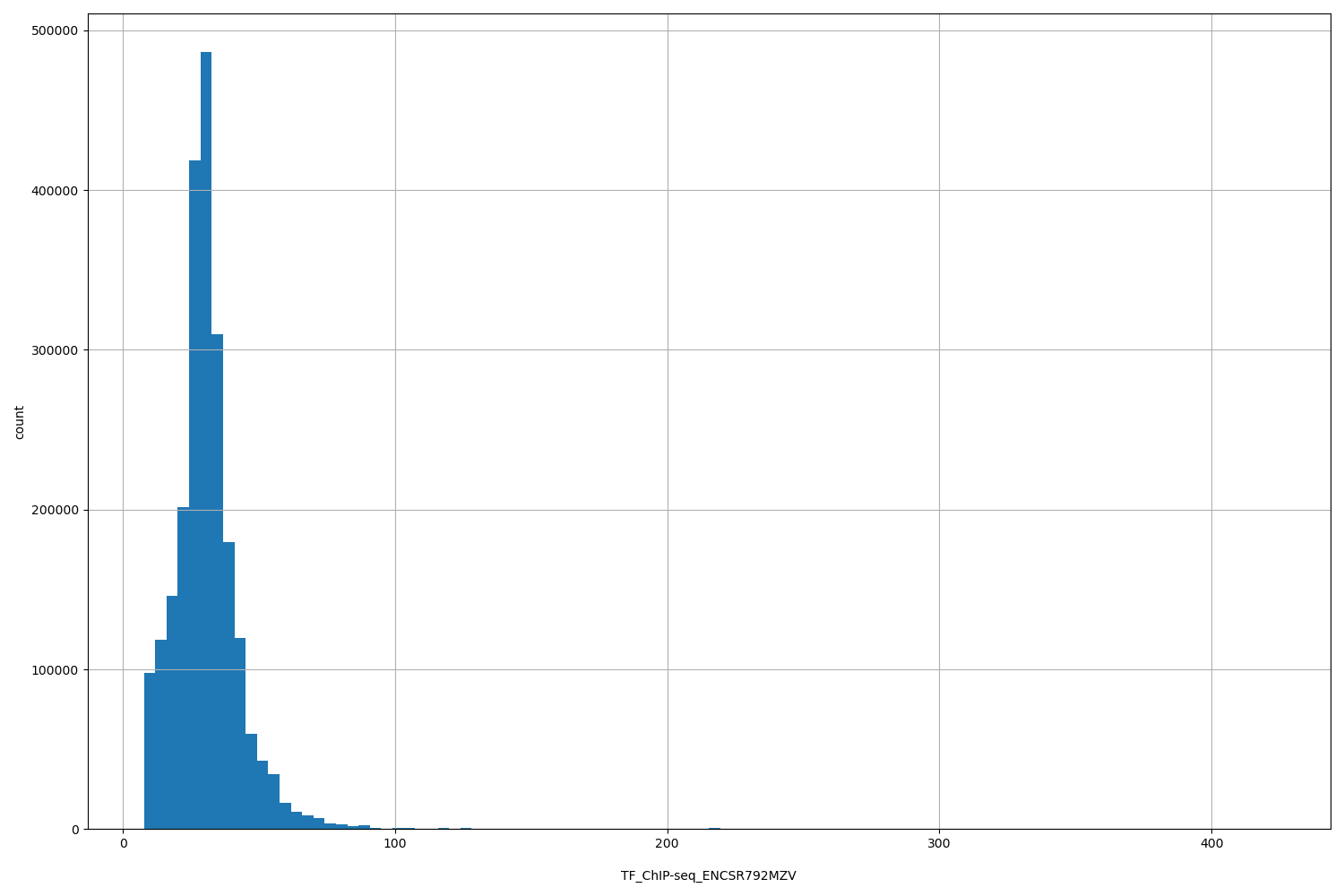
TF_ChIP-seq_ENCSR792MZV
| Id: | TF_ChIP-seq/ENCSR792MZV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR792MZV [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens MYRF" and target="MYRF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens MYRF output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN740DHN|/analyses/ENCAN740DHN/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF800FXA|/files/ENCFF800FXA/}, {ENCFF335DMN|/files/ENCFF335DMN/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.41 and a self consistency ratio of 2.07. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF650LIT|/files/ENCFF650LIT/}, {ENCFF804RUY|/files/ENCFF804RUY/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.41 and a self consistency ratio of 2.07. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR792MZV | float |
TF_ChIP-seq_ENCSR792MZV |
TF_ChIP-seq ENCSR792MZV [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens MYRF" and target="MYRF"]
|
 |
[7.73, 423] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF804RUY.bed.gz | 123.0 KB | 8919853879d9fd8847317a9f728a4010 |
| ENCFF804RUY.bed.gz.dvc | 100.0 B | 79640d6cdf30980187b5f41171983cbe |
| ENCFF804RUY.tabix.bed.gz | 86.6 KB | 89e68e8fc9ca9267dd10f453a62a86e8 |
| ENCFF804RUY.tabix.bed.gz.dvc | 105.0 B | 0cfe536d363e91384ab48c7e43198e50 |
| ENCFF804RUY.tabix.bed.gz.tbi | 51.84 KB | a512a2dfc0e91403adacec49d4b824e8 |
| ENCFF804RUY.tabix.bed.gz.tbi.dvc | 109.0 B | 3531029b15a5da8a2e363c4be48a6fd2 |
| genomic_resource.yaml | 2.79 KB | 6f06d2e2944eba726f6c8d846d023bfe |
| genomic_resource_original.yaml | 2.63 KB | 06cef2834d60699b6dec8c26aefdcb13 |
| statistics/ |