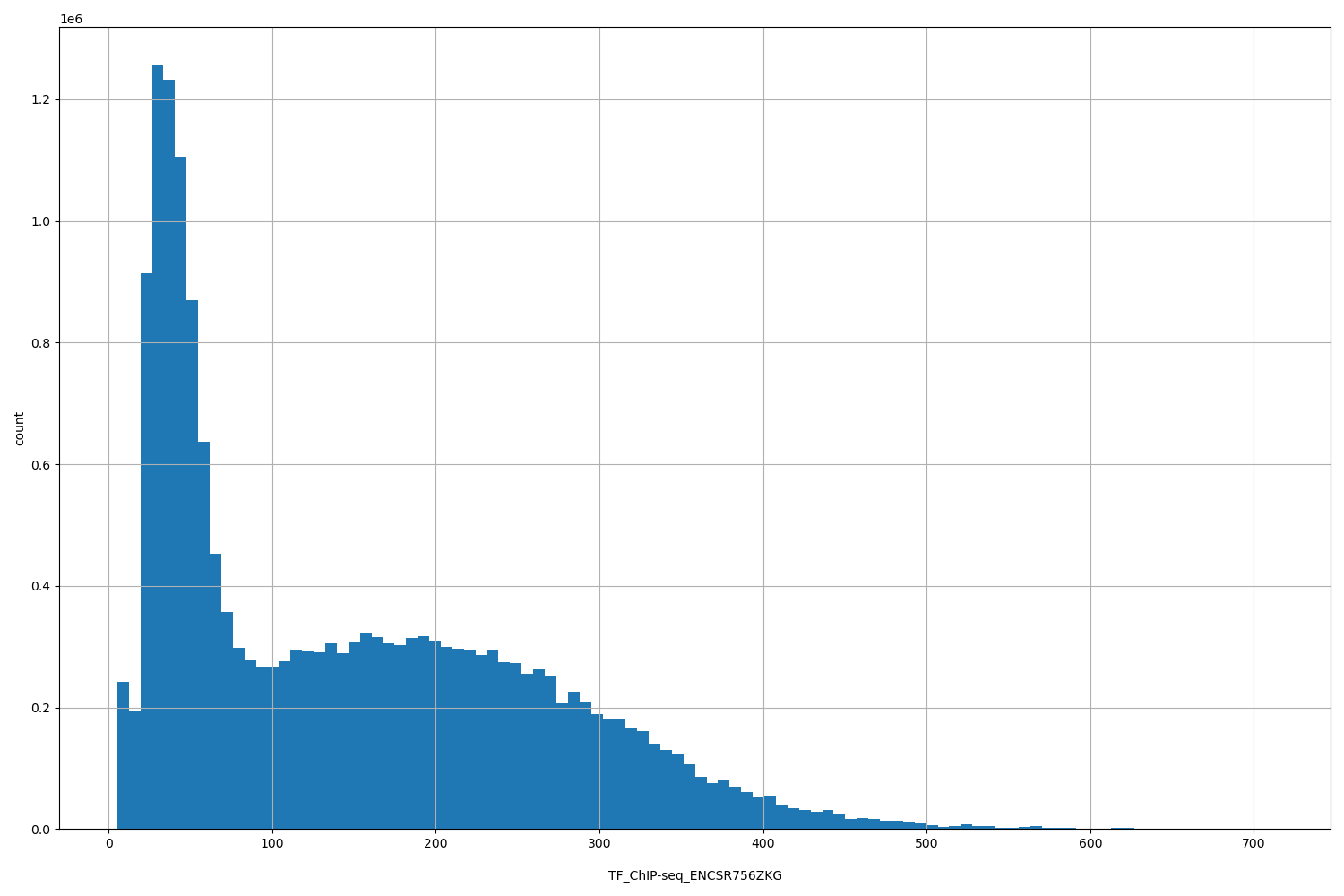
TF_ChIP-seq_ENCSR756ZKG
| Id: | TF_ChIP-seq/ENCSR756ZKG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR756ZKG [biosamplesummary="Homo sapiens OCI-LY3" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN833OGF|/analyses/ENCAN833OGF/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF496TSK|/files/ENCFF496TSK/} processed by ChIP-seq ENCODE3 hg19 pipeline has 16509737 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR756ZKG | float |
TF_ChIP-seq_ENCSR756ZKG |
TF_ChIP-seq ENCSR756ZKG [biosample_summary="Homo sapiens OCI-LY3" and target="CTCF"]
|
 |
[5.26, 712] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF653ZMJ.bed.gz | 913.13 KB | 39fc52436e5ef8b2ecdc550dde48f11d |
| ENCFF653ZMJ.bed.gz.dvc | 100.0 B | 497ef2a63731e4f78ac96e79fdaf6423 |
| ENCFF653ZMJ.tabix.bed.gz | 684.17 KB | 13aa7314cbb03771d58bdc68552d15ef |
| ENCFF653ZMJ.tabix.bed.gz.dvc | 106.0 B | a769f64052165fe34345502d11840f14 |
| ENCFF653ZMJ.tabix.bed.gz.tbi | 329.12 KB | 50ed72d044459bb5a12732ad1df3af67 |
| ENCFF653ZMJ.tabix.bed.gz.tbi.dvc | 110.0 B | b533f42cc0f9b570304e3c24d545f1ac |
| genomic_resource.yaml | 1.84 KB | 431e55af38756f123f027aab33a65224 |
| genomic_resource_original.yaml | 1.74 KB | 9c6dc0a8666075480816990ee511a8f0 |
| statistics/ |