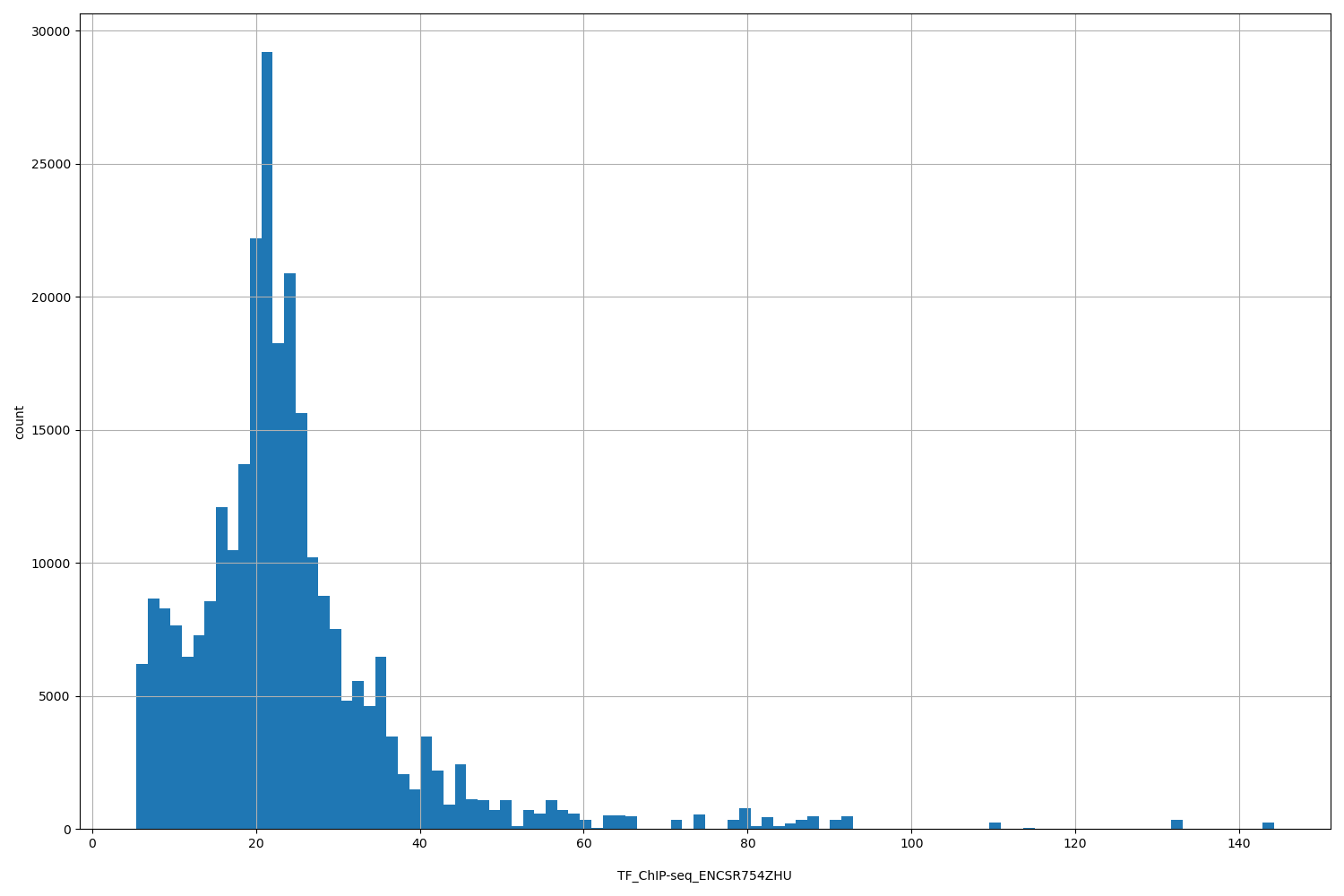
TF_ChIP-seq_ENCSR754ZHU
| Id: | TF_ChIP-seq/ENCSR754ZHU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR754ZHU [biosamplesummary="Homo sapiens K562" and target="SNRNP70"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: conservative IDR thresholded peaks audit_internal_action: Released analysis {ENCAN226GEQ|/analyses/ENCAN226GEQ/} has in progress subobject document {2b05ec5b-e768-4b34-9a42-cebd0691c8c8|/documents/2b05ec5b-e768-4b34-9a42-cebd0691c8c8/} audit_internal_action: Released analysis {ENCAN226GEQ|/analyses/ENCAN226GEQ/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF507MYK|/files/ENCFF507MYK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 15643332 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SNRNP70-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF570NLH|/files/ENCFF570NLH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16681103 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SNRNP70-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR754ZHU | float |
TF_ChIP-seq_ENCSR754ZHU |
TF_ChIP-seq ENCSR754ZHU [biosample_summary="Homo sapiens K562" and target="SNRNP70"]
|
 |
[5.41, 144] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF306TAD.bed.gz | 15.72 KB | b7bccec84c390a27a5402d2224b8a277 |
| ENCFF306TAD.bed.gz.dvc | 99.0 B | 85690629419a8bb3e47c457dddaef170 |
| ENCFF306TAD.tabix.bed.gz | 10.88 KB | a9ee8cc5eb186ceb91be165455865018 |
| ENCFF306TAD.tabix.bed.gz.dvc | 105.0 B | 881c068298ada46cbabf51ee06a6ea4f |
| ENCFF306TAD.tabix.bed.gz.tbi | 9.44 KB | c638843b6da297251b2c2f1261c87e9a |
| ENCFF306TAD.tabix.bed.gz.tbi.dvc | 108.0 B | 0e6fbb640b27fb312763d4bf332c79e3 |
| genomic_resource.yaml | 2.58 KB | daaa4ca8ffb7fdc3608929bf517e9452 |
| genomic_resource_original.yaml | 2.48 KB | 7560d7c12baed93d0162f6bb14b29db6 |
| statistics/ |