TF_ChIP-seq_ENCSR736BUG
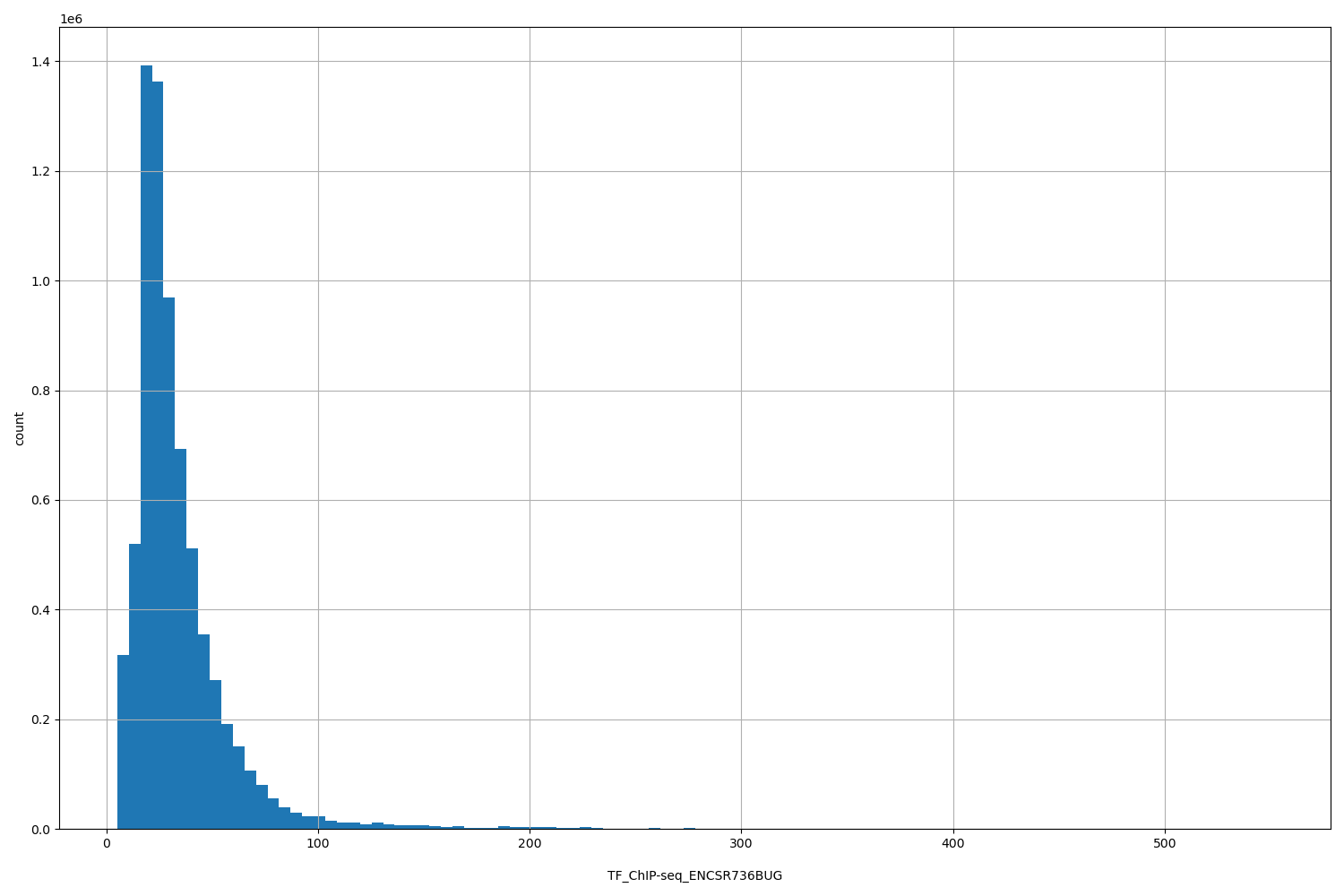
| Id: | TF_ChIP-seq/ENCSR736BUG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR736BUG [biosamplesummary="Homo sapiens with nonobstructive coronary artery disease; liver tissue male adult (32 years)" and target="EGR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (32 years) with nonobstructive coronary artery disease output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF617JQS|/files/ENCFF617JQS/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN782ICQ|/analyses/ENCAN782ICQ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF174ZGG|/files/ENCFF174ZGG/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18205232 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EGR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF995SEN|/files/ENCFF995SEN/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 17605252 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EGR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR736BUG | float |
TF_ChIP-seq_ENCSR736BUG |
TF_ChIP-seq ENCSR736BUG [biosample_summary="Homo sapiens with nonobstructive coronary artery disease; liver tissue male adult (32 years)" and target="EGR1"]
|
 |
[5.13, 551] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF617JQS.bed.gz | 333.91 KB | 49b41dece0ca596f3d0edc8cec227e0f |
| ENCFF617JQS.bed.gz.dvc | 100.0 B | df6909a5fd1bb0e590975b6f45f297d4 |
| ENCFF617JQS.tabix.bed.gz | 253.34 KB | 5abc8a052917ebc0be4ace90506c2b6d |
| ENCFF617JQS.tabix.bed.gz.dvc | 106.0 B | ede3d262119b7384a9b555427e11e6d5 |
| ENCFF617JQS.tabix.bed.gz.tbi | 125.98 KB | 42c98fe9800751874707c456bf66b897 |
| ENCFF617JQS.tabix.bed.gz.tbi.dvc | 110.0 B | d2c34481417b11715631ca51a4b9da82 |
| genomic_resource.yaml | 3.17 KB | 4759feafd4769b878ee0b62ac81fe73f |
| genomic_resource_original.yaml | 3.01 KB | 6af611cfebb36d9f2f0938307b6352f7 |
| statistics/ |