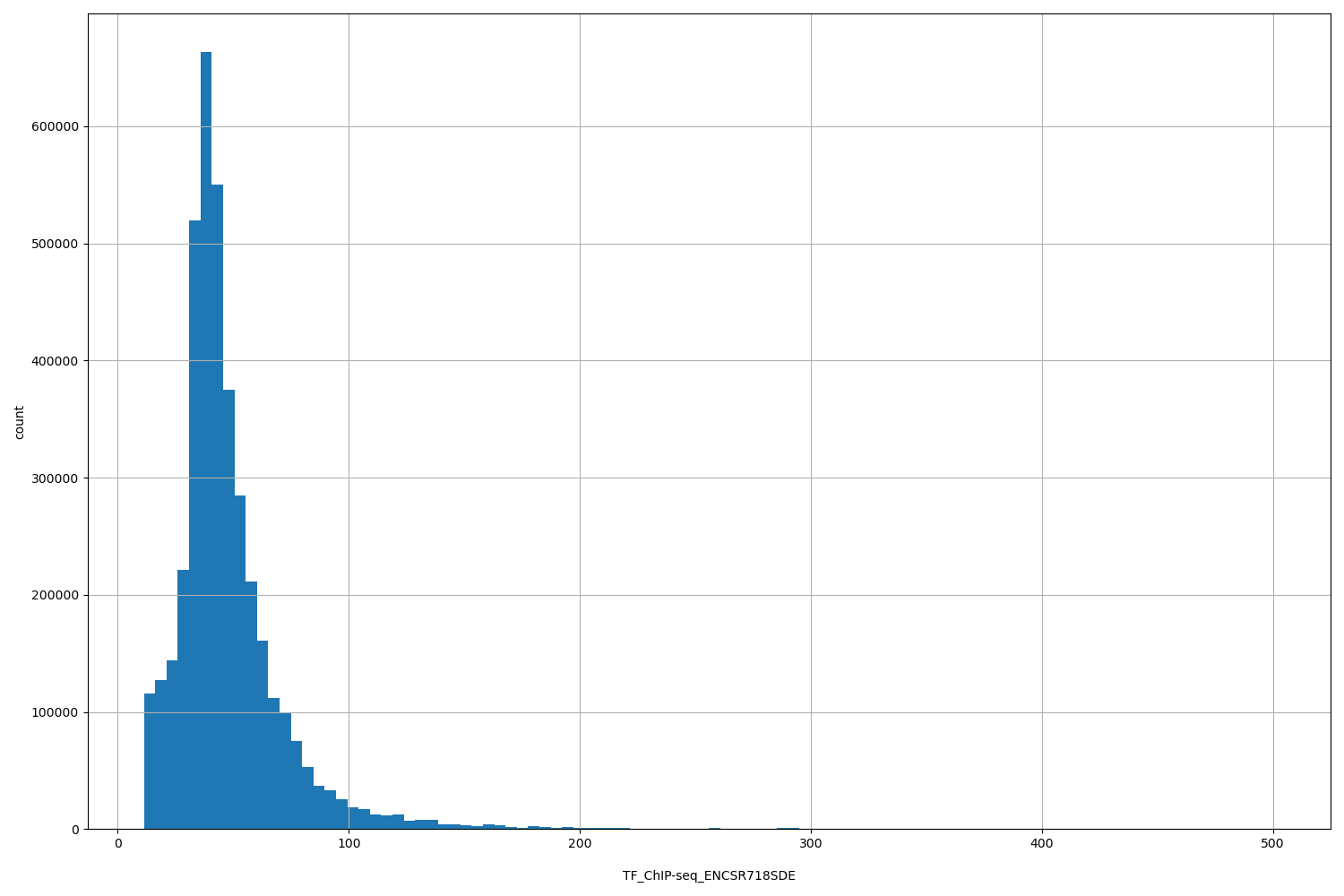
TF_ChIP-seq_ENCSR718SDE
| Id: | TF_ChIP-seq/ENCSR718SDE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR718SDE [biosamplesummary="Homo sapiens K562" and target="RLF"] |
| Description: |
status: released biological_replicates: Rep 2, Rep 3 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN484VAX|/analyses/ENCAN484VAX/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF243VKO|/files/ENCFF243VKO/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19647918 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting RLF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR718SDE | float |
TF_ChIP-seq_ENCSR718SDE |
TF_ChIP-seq ENCSR718SDE [biosample_summary="Homo sapiens K562" and target="RLF"]
|
 |
[11.3, 500] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF209HKW.bed.gz | 195.88 KB | 0715e48bb3bd6c77a51ee4cda8cb0a97 |
| ENCFF209HKW.bed.gz.dvc | 100.0 B | e46a8952f4c65e48122d506787d852ab |
| ENCFF209HKW.tabix.bed.gz | 137.12 KB | 1a9ee52fc72cf64da11f09547471bd30 |
| ENCFF209HKW.tabix.bed.gz.dvc | 106.0 B | e41f12e29c6c01cf6f8246106de7391a |
| ENCFF209HKW.tabix.bed.gz.tbi | 74.61 KB | 7e1c73a44841aae5e25b3d4b7c3ebc05 |
| ENCFF209HKW.tabix.bed.gz.tbi.dvc | 109.0 B | fda9dd14916bfea4252a707b7965030e |
| genomic_resource.yaml | 2.37 KB | c2b081e8e03949f0f3b1cd3e59403b7e |
| genomic_resource_original.yaml | 2.28 KB | f7398f2b4d7c7921672e6e52108f8ab5 |
| statistics/ |