TF_ChIP-seq_ENCSR707BNG
| Id: | TF_ChIP-seq/ENCSR707BNG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR707BNG [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens MYNN" and target="MYNN"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens MYNN output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN964FET|/analyses/ENCAN964FET/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF786FAF|/files/ENCFF786FAF/}, {ENCFF045LTC|/files/ENCFF045LTC/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.59 and a self consistency ratio of 2.33. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF579EPP|/files/ENCFF579EPP/}, {ENCFF653XGP|/files/ENCFF653XGP/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.59 and a self consistency ratio of 2.33. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
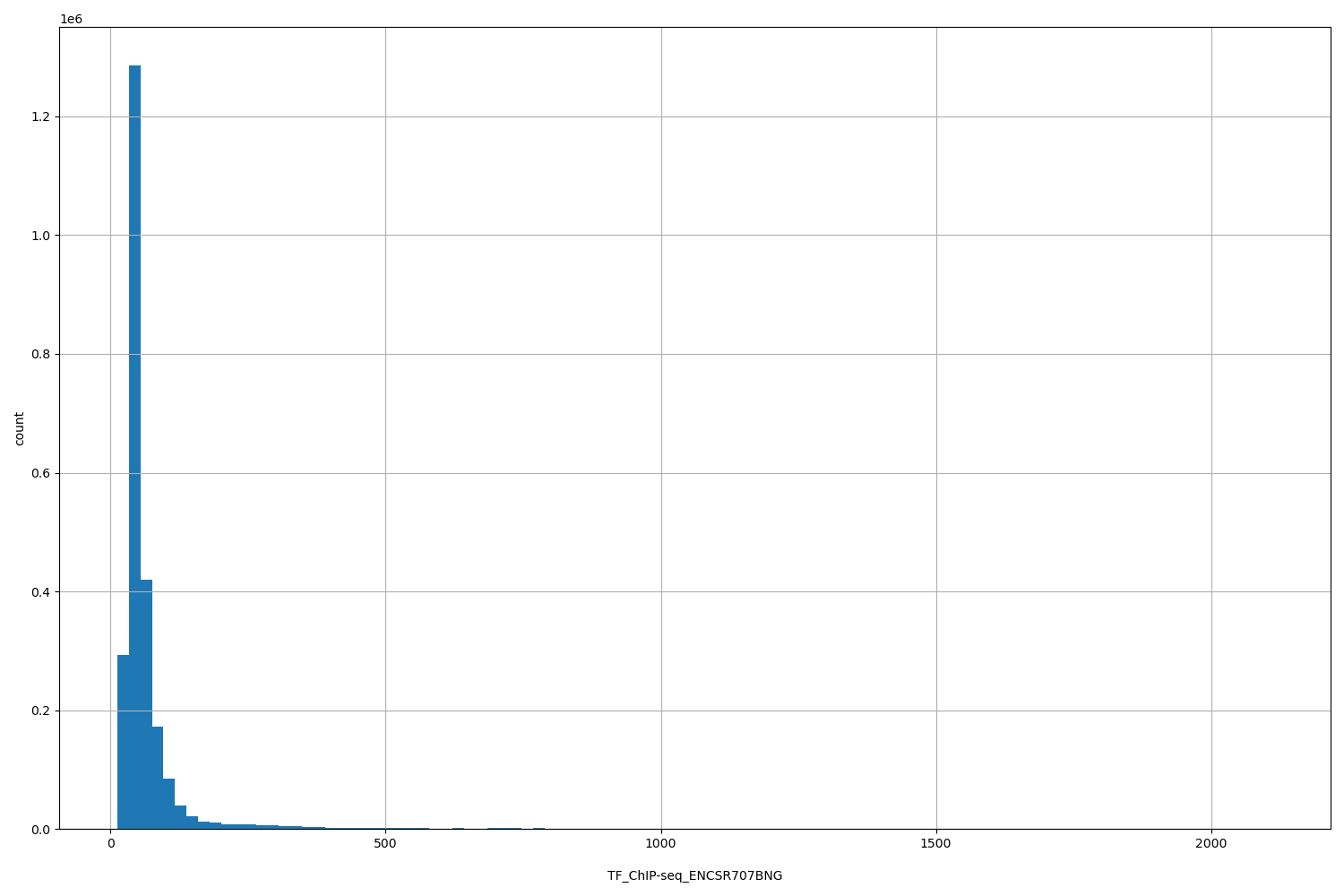
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR707BNG | float |
TF_ChIP-seq_ENCSR707BNG |
TF_ChIP-seq ENCSR707BNG [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens MYNN" and target="MYNN"]
|
 |
[12.4, 2.11e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF045LTC.bed.gz | 129.95 KB | 78e40931e1277871066a6dd063f55dca |
| ENCFF045LTC.bed.gz.dvc | 100.0 B | 4579d833d5dfa1a8e2261f139302845c |
| ENCFF045LTC.tabix.bed.gz | 88.51 KB | d3e8eaa3598271bd91295dd25e3334ef |
| ENCFF045LTC.tabix.bed.gz.dvc | 105.0 B | 40f12a5d7eae0cbbc4ee4140eb370a4a |
| ENCFF045LTC.tabix.bed.gz.tbi | 59.36 KB | 8b82f72cd0f7b10c5edb47bcf88462d4 |
| ENCFF045LTC.tabix.bed.gz.tbi.dvc | 109.0 B | 07c4ce7d263599bad8f770f91cf85eee |
| genomic_resource.yaml | 2.9 KB | b2f2c48609e8f9d04270754e445c8a22 |
| genomic_resource_original.yaml | 2.72 KB | 64087455eed3f8878e149cb589070e27 |
| statistics/ |