TF_ChIP-seq_ENCSR666QNP
| Id: | TF_ChIP-seq/ENCSR666QNP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR666QNP [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens TEAD3" and target="TEAD3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens TEAD3 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN628OCL|/analyses/ENCAN628OCL/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF802TPU|/files/ENCFF802TPU/}, {ENCFF257QVH|/files/ENCFF257QVH/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.42 and a self consistency ratio of 2.54. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF138XKG|/files/ENCFF138XKG/}, {ENCFF079MKQ|/files/ENCFF079MKQ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.42 and a self consistency ratio of 2.54. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
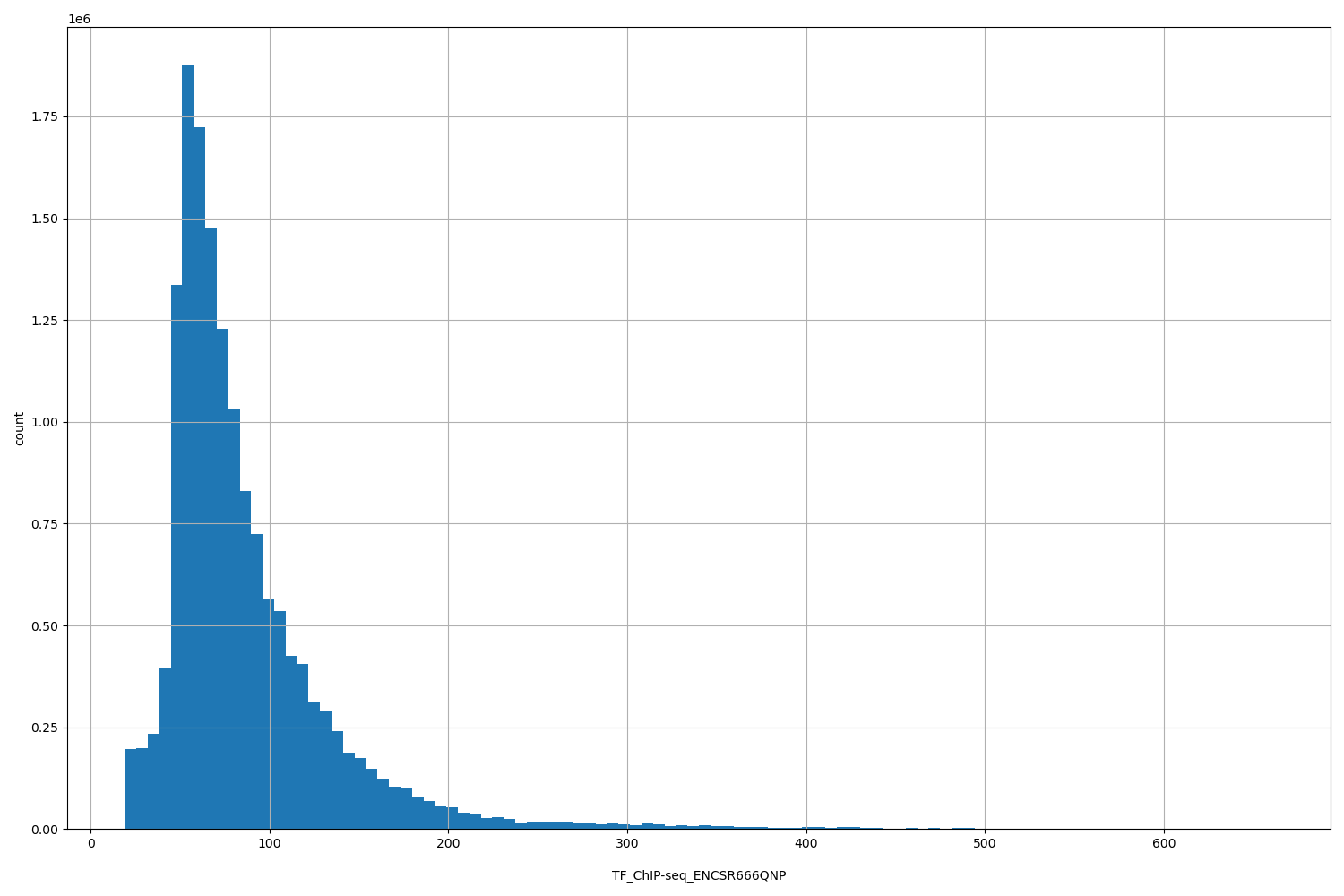
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR666QNP | float |
TF_ChIP-seq_ENCSR666QNP |
TF_ChIP-seq ENCSR666QNP [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens TEAD3" and target="TEAD3"]
|
 |
[19.1, 661] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF079MKQ.bed.gz | 772.89 KB | a034665246745316d8f1db2a5fd668db |
| ENCFF079MKQ.bed.gz.dvc | 100.0 B | 04fcd13eeedf1d42a96c777b71679e98 |
| ENCFF079MKQ.tabix.bed.gz | 557.93 KB | 564be5f6d9f89f87cf3bfcf534dad260 |
| ENCFF079MKQ.tabix.bed.gz.dvc | 106.0 B | 0eedbd150b41e0d7b258754fb61b05e1 |
| ENCFF079MKQ.tabix.bed.gz.tbi | 222.27 KB | 6893a5eca99362a6fba5d7099156ee52 |
| ENCFF079MKQ.tabix.bed.gz.tbi.dvc | 110.0 B | e83f1260130a615ae73a27a513b11b3c |
| genomic_resource.yaml | 2.86 KB | 6676030ae3ef65c22cb959cf4e0aba81 |
| genomic_resource_original.yaml | 2.69 KB | b83437f98fee7cbb4fd2bf53aa4f61f8 |
| statistics/ |