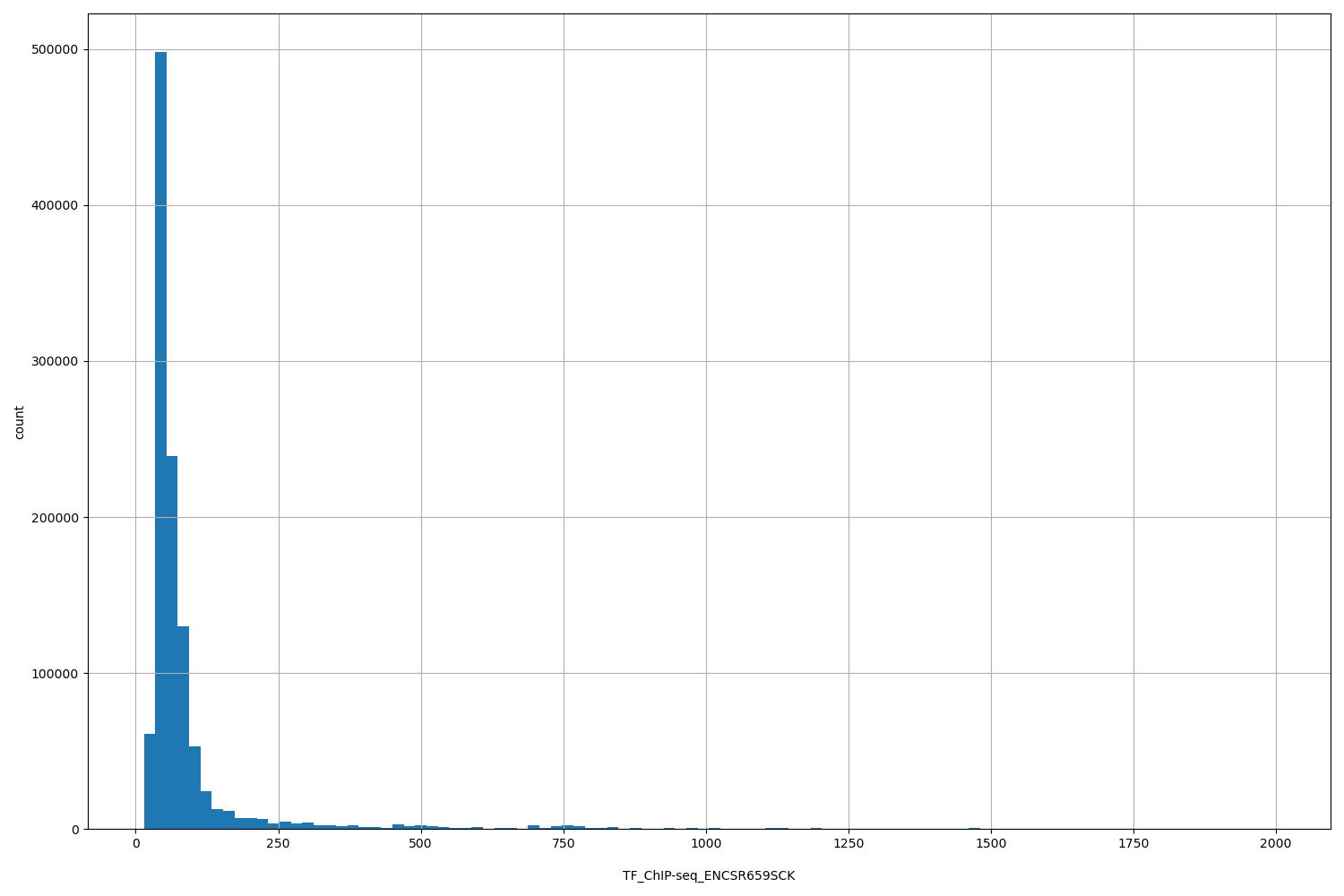
TF_ChIP-seq_ENCSR659SCK
| Id: | TF_ChIP-seq/ENCSR659SCK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR659SCK [biosamplesummary="Homo sapiens HepG2" and target="TOE1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN612ANU|/analyses/ENCAN612ANU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF539FVQ|/files/ENCFF539FVQ/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF117NTL|/files/ENCFF117NTL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.80. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF117NTL|/files/ENCFF117NTL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 4.95. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF310XQA|/files/ENCFF310XQA/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.86. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF310XQA|/files/ENCFF310XQA/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 7.10. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF862TRT|/files/ENCFF862TRT/}, {ENCFF574HTD|/files/ENCFF574HTD/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.60 and a self consistency ratio of 2.90. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF539FVQ|/files/ENCFF539FVQ/}, {ENCFF801JVM|/files/ENCFF801JVM/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.60 and a self consistency ratio of 2.90. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR659SCK | float |
TF_ChIP-seq_ENCSR659SCK |
TF_ChIP-seq ENCSR659SCK [biosample_summary="Homo sapiens HepG2" and target="TOE1"]
|
 |
[14.3, 2e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF539FVQ.bed.gz | 74.02 KB | 0341078fee5d5acf8858826e469dfb0d |
| ENCFF539FVQ.bed.gz.dvc | 99.0 B | f981eda4e998bf4a99b308c00737f311 |
| ENCFF539FVQ.tabix.bed.gz | 49.15 KB | 9a2fab0438259a2318b8672b97065034 |
| ENCFF539FVQ.tabix.bed.gz.dvc | 105.0 B | a99afababcfa7d2a917970238ea3d3e0 |
| ENCFF539FVQ.tabix.bed.gz.tbi | 43.2 KB | a149bc31aa56a88ddf5424237c874fee |
| ENCFF539FVQ.tabix.bed.gz.tbi.dvc | 109.0 B | 2a6c1d57d17ad56ccd2498c0c11a1973 |
| genomic_resource.yaml | 5.26 KB | ec895338fad0c84590dea70dba3af8ba |
| genomic_resource_original.yaml | 5.17 KB | 8307a26050cea7673682e5298192347f |
| statistics/ |