TF_ChIP-seq_ENCSR626VUC
| Id: | TF_ChIP-seq/ENCSR626VUC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR626VUC [biosamplesummary="Homo sapiens GM12878" and target="ETV6"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF745ANU|/files/ENCFF745ANU/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Released file {ENCFF745ANU|/files/ENCFF745ANU/} has replaced subobject file {a4bf1d72-be83-42fc-95ca-04d9a344600a|/files/a4bf1d72-be83-42fc-95ca-04d9a344600a/} audit_internal_action: Archived analysis {ENCAN809OOT|/analyses/ENCAN809OOT/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF269LTR|/files/ENCFF269LTR/}, {ENCFF745ANU|/files/ENCFF745ANU/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.50 and a self consistency ratio of 2.39. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF592LKK|/files/ENCFF592LKK/}, {ENCFF482BSZ|/files/ENCFF482BSZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.50 and a self consistency ratio of 2.39. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
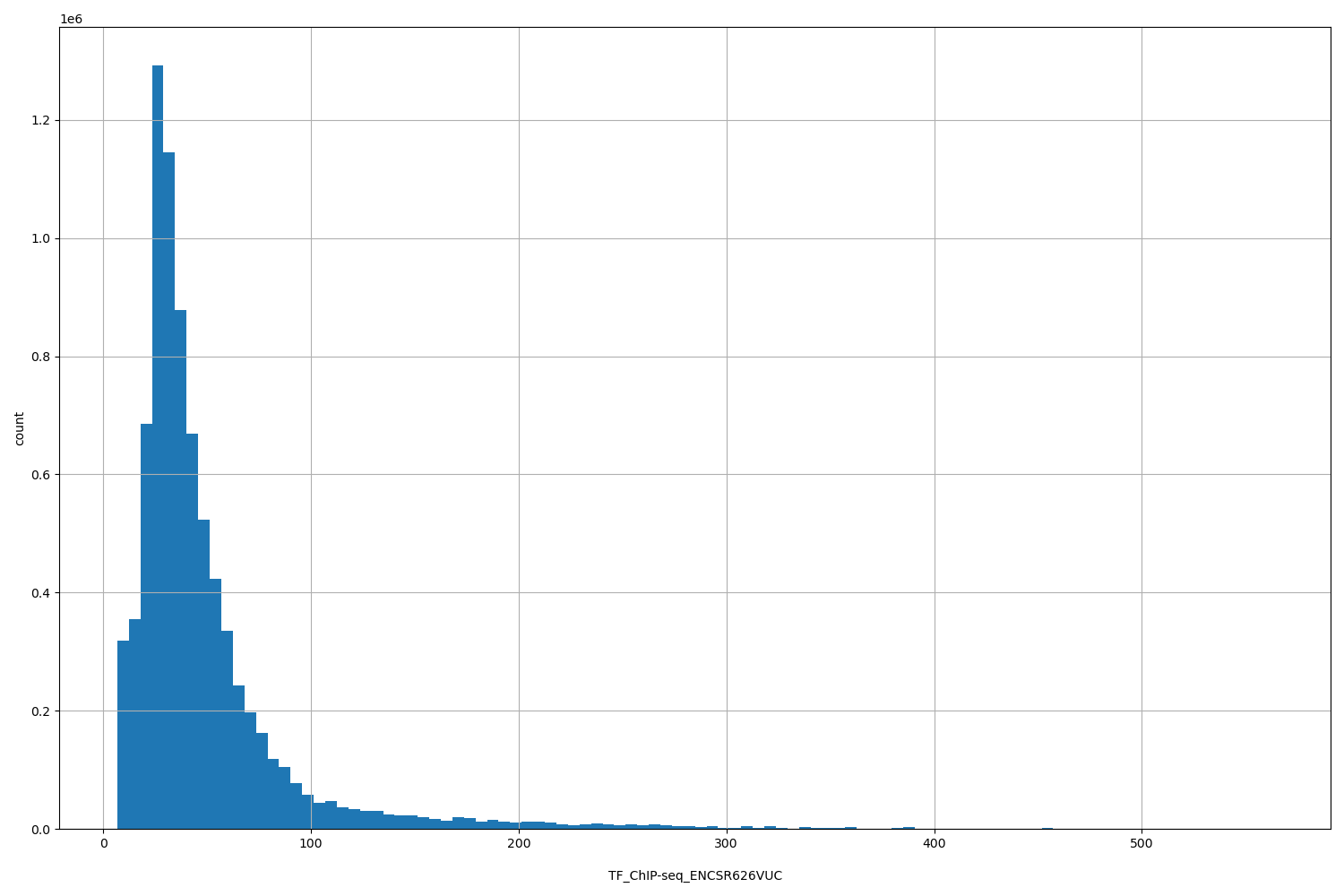
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR626VUC | float |
TF_ChIP-seq_ENCSR626VUC |
TF_ChIP-seq ENCSR626VUC [biosample_summary="Homo sapiens GM12878" and target="ETV6"]
|
 |
[6.71, 563] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF745ANU.bed.gz | 373.07 KB | cff6f88f836c9a4566acf3a4bc39d9a0 |
| ENCFF745ANU.bed.gz.dvc | 100.0 B | 3522114606eb025c544808eadcab8efe |
| ENCFF745ANU.tabix.bed.gz | 290.06 KB | 60a6c90b437be37efdb0f00e00fe076b |
| ENCFF745ANU.tabix.bed.gz.dvc | 106.0 B | 0a1712a18ab6ecc021caa971d0622219 |
| ENCFF745ANU.tabix.bed.gz.tbi | 126.86 KB | ea094e637844545f21a3a0c262c942ef |
| ENCFF745ANU.tabix.bed.gz.tbi.dvc | 110.0 B | 5d15e5a4eb1542a41c2a9fd1e4a434ee |
| genomic_resource.yaml | 2.87 KB | 82608e6ff76bd83d9e76358d23bf43df |
| genomic_resource_original.yaml | 2.77 KB | 24ac07b8efd0f00b2e6278269ffc7dea |
| statistics/ |