TF_ChIP-seq_ENCSR616WEG
| Id: | TF_ChIP-seq/ENCSR616WEG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR616WEG [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HBP1" and target="HBP1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens HBP1 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN512FCJ|/analyses/ENCAN512FCJ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF998QTY|/files/ENCFF998QTY/} processed by ChIP-seq ENCODE3 hg19 pipeline has 16223700 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting HBP1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF431UUS|/files/ENCFF431UUS/} processed by ChIP-seq ENCODE3 hg19 pipeline has 16732372 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting HBP1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
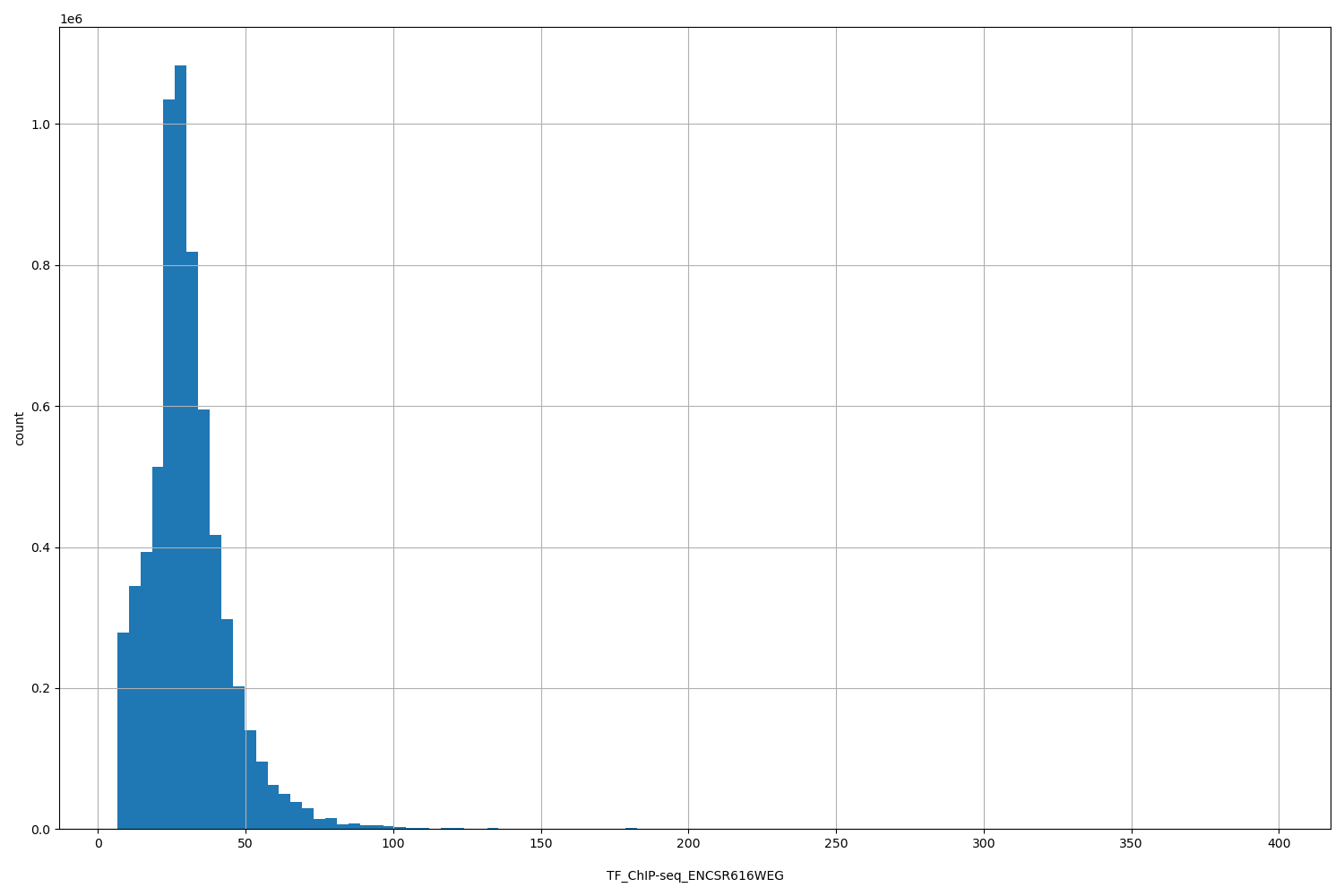
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR616WEG | float |
TF_ChIP-seq_ENCSR616WEG |
TF_ChIP-seq ENCSR616WEG [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HBP1" and target="HBP1"]
|
 |
[6.5, 398] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF104YWL.bed.gz | 270.95 KB | a50703f33fc1527de3069617ddc8f8e1 |
| ENCFF104YWL.bed.gz.dvc | 100.0 B | d7d013db4df2ebb3a4f843b93356b550 |
| ENCFF104YWL.tabix.bed.gz | 211.02 KB | 003926bff6e38142b9f1eeef477c0b66 |
| ENCFF104YWL.tabix.bed.gz.dvc | 106.0 B | 54f37163900247fb3a4c24bcc22f786f |
| ENCFF104YWL.tabix.bed.gz.tbi | 96.75 KB | c99f5a2caea8e8c53f781cc77bcd7fa0 |
| ENCFF104YWL.tabix.bed.gz.tbi.dvc | 109.0 B | 00e58f41a95cf1a38e4c1022f638b471 |
| genomic_resource.yaml | 2.65 KB | f0e138ba9f60f93a9cab57f0ac2706eb |
| genomic_resource_original.yaml | 2.49 KB | 9a53b3cf3488d4c23e60875ae9b9c5fa |
| statistics/ |