TF_ChIP-seq_ENCSR608XTF
| Id: | TF_ChIP-seq/ENCSR608XTF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR608XTF [biosamplesummary="Homo sapiens K562" and target="RNF2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN003ACX|/analyses/ENCAN003ACX/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF462AZY|/files/ENCFF462AZY/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF779IQL|/files/ENCFF779IQL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.83. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF779IQL|/files/ENCFF779IQL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 5.84. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF257OOQ|/files/ENCFF257OOQ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.83. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF257OOQ|/files/ENCFF257OOQ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 5.80. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF462AZY|/files/ENCFF462AZY/}, {ENCFF389LKZ|/files/ENCFF389LKZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 11.19 and a self consistency ratio of 1.64. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF239KJX|/files/ENCFF239KJX/}, {ENCFF173PIB|/files/ENCFF173PIB/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 11.19 and a self consistency ratio of 1.64. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
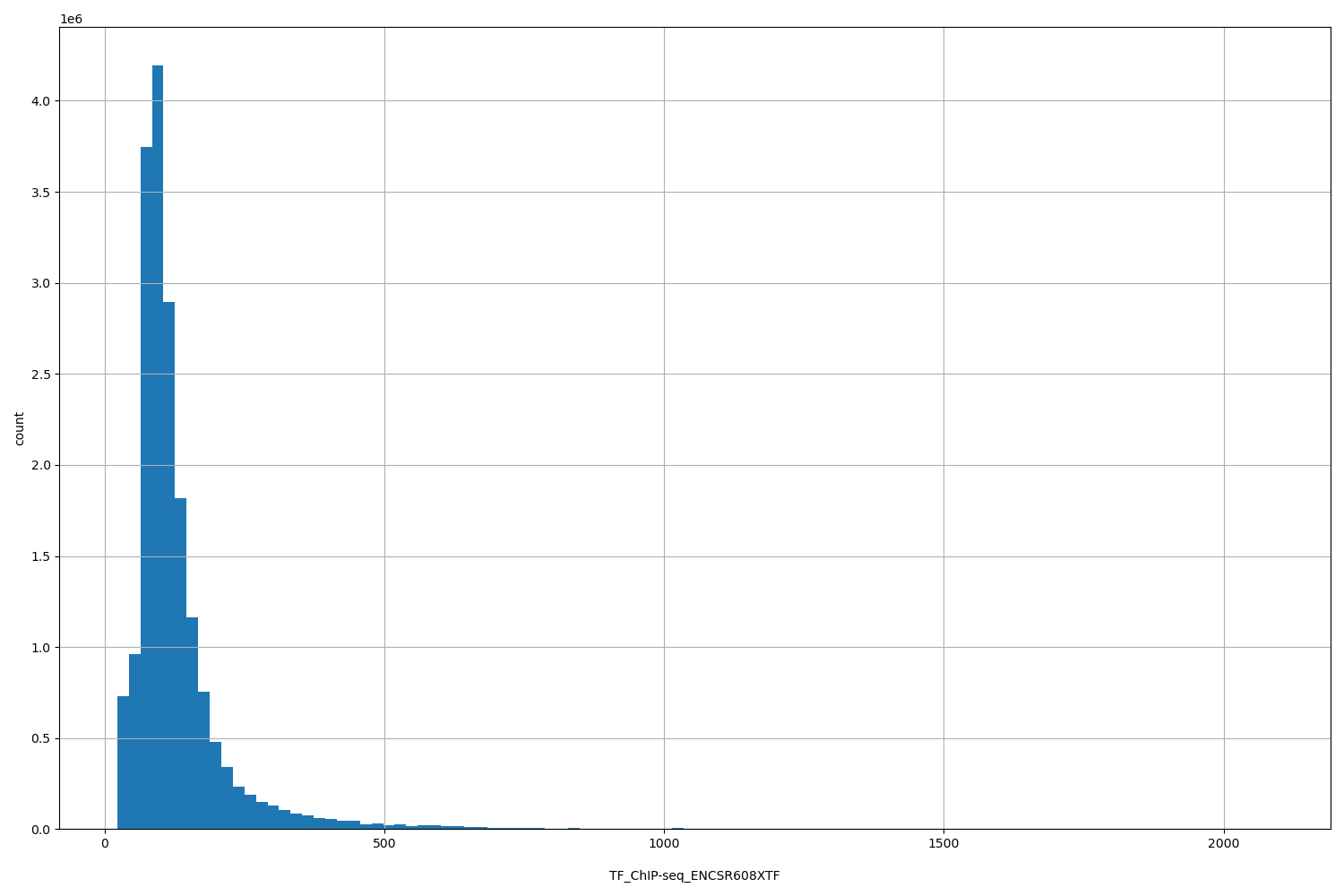
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR608XTF | float |
TF_ChIP-seq_ENCSR608XTF |
TF_ChIP-seq ENCSR608XTF [biosample_summary="Homo sapiens K562" and target="RNF2"]
|
 |
[23, 2.09e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF462AZY.bed.gz | 716.69 KB | 5a66428f408e8eb4593f0ec2663cebff |
| ENCFF462AZY.bed.gz.dvc | 100.0 B | 686cac1a6d4f08a89e1de898ce0459ff |
| ENCFF462AZY.tabix.bed.gz | 513.53 KB | 4a6884ac921dbc436606d7be2185b51a |
| ENCFF462AZY.tabix.bed.gz.dvc | 106.0 B | 656f921e7a63fa7ef300ba1894a52b82 |
| ENCFF462AZY.tabix.bed.gz.tbi | 190.54 KB | 63213c4e3841d3aa20258442345bffd1 |
| ENCFF462AZY.tabix.bed.gz.tbi.dvc | 110.0 B | 0f008b6da07240362df37debeb8ca28b |
| genomic_resource.yaml | 5.26 KB | 6c18e1b305fa52a73683dfed63234fed |
| genomic_resource_original.yaml | 5.17 KB | f7d3a64212a781b5be8aaaf09378edc6 |
| statistics/ |