TF_ChIP-seq_ENCSR580IAO
| Id: | TF_ChIP-seq/ENCSR580IAO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR580IAO [biosamplesummary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF197" and target="ZNF197"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZNF197 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN645PAT|/analyses/ENCAN645PAT/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF764PZX|/files/ENCFF764PZX/} processed by ChIP-seq ENCODE3 hg19 pipeline has 14788885 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF197-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF947CVN|/files/ENCFF947CVN/} processed by ChIP-seq ENCODE3 hg19 pipeline has 13738772 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF197-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
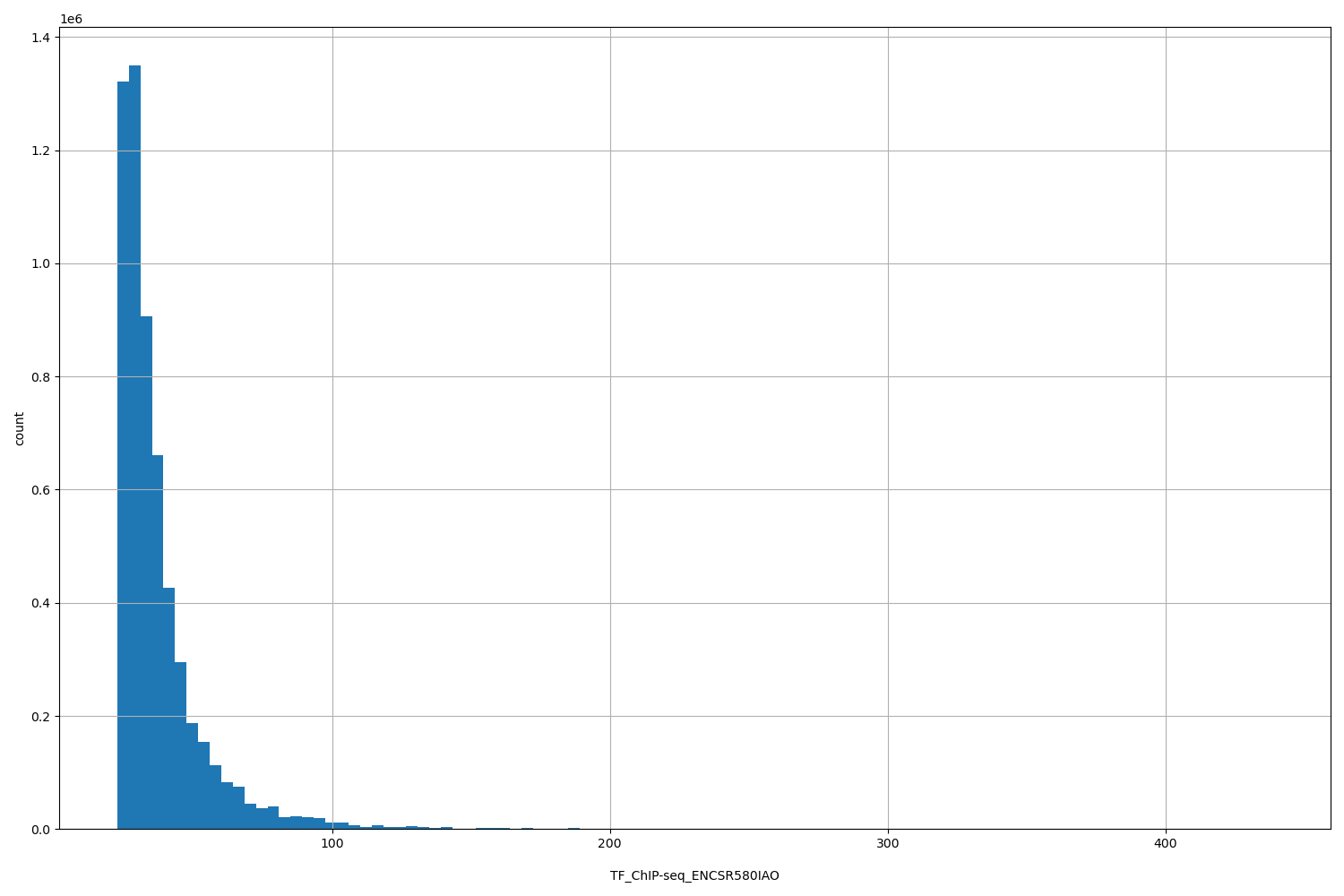
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR580IAO | float |
TF_ChIP-seq_ENCSR580IAO |
TF_ChIP-seq ENCSR580IAO [biosample_summary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF197" and target="ZNF197"]
|
 |
[22.7, 439] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF853TZC.bed.gz | 157.87 KB | 76dff59260974ea4959eabcd3f275a6a |
| ENCFF853TZC.bed.gz.dvc | 100.0 B | 4ffc3ea68367b61f24bfd68cecd25316 |
| ENCFF853TZC.tabix.bed.gz | 145.05 KB | 46a0090442c2edf46027a91431307415 |
| ENCFF853TZC.tabix.bed.gz.dvc | 106.0 B | 17e54de0efbc3af12dfa58183ee72669 |
| ENCFF853TZC.tabix.bed.gz.tbi | 67.51 KB | 505a1492ab84d7f9fbc6d7b9edb19d2c |
| ENCFF853TZC.tabix.bed.gz.tbi.dvc | 109.0 B | d550a5c354cdba8a63b6ba2afd091274 |
| genomic_resource.yaml | 2.66 KB | b8e0cea71b8d1029d5e61fe605b7cecd |
| genomic_resource_original.yaml | 2.5 KB | b1ba38885bb8c27c8c2fa0f28fc67c82 |
| statistics/ |