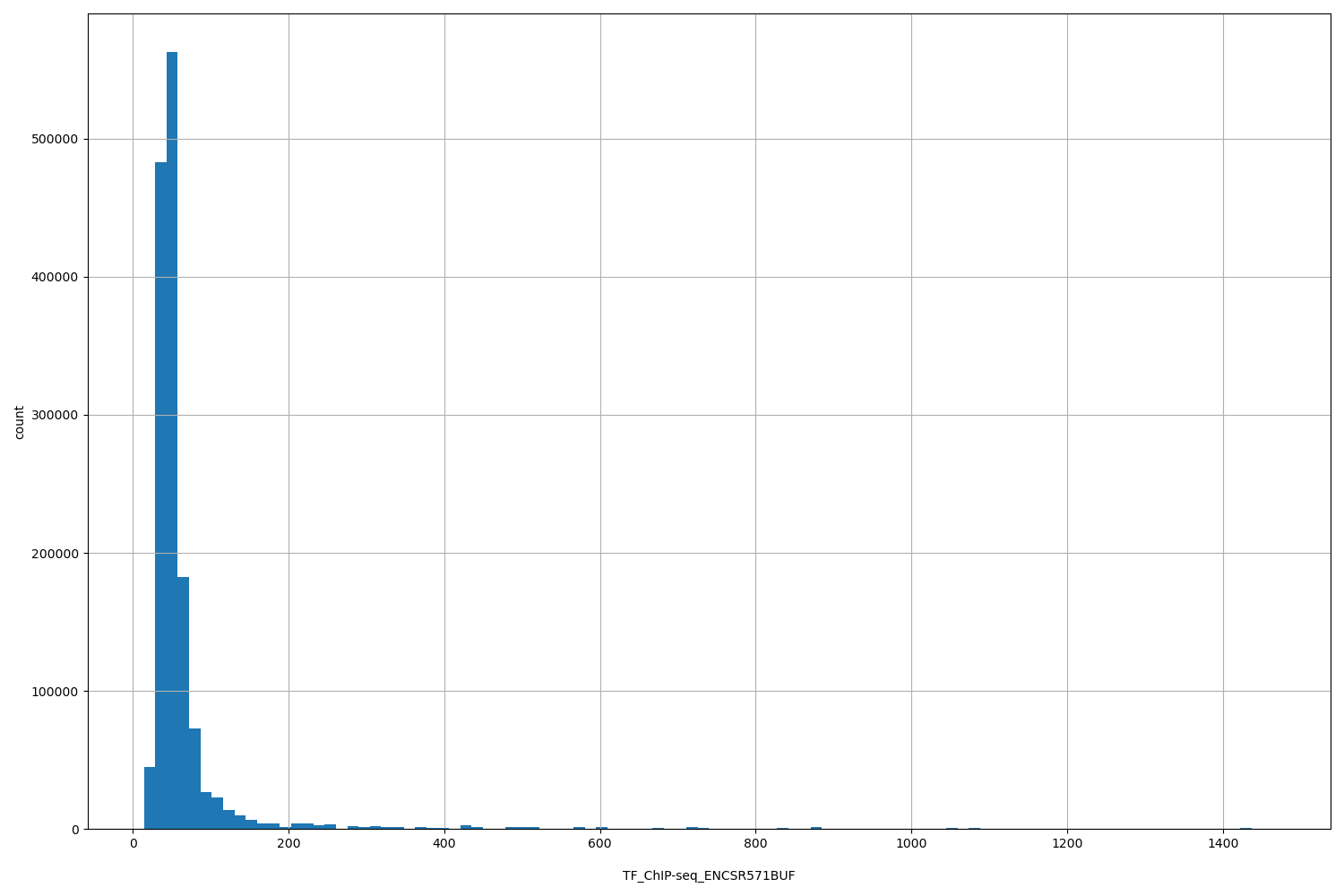
TF_ChIP-seq_ENCSR571BUF
| Id: | TF_ChIP-seq/ENCSR571BUF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR571BUF [biosamplesummary="Homo sapiens K562" and target="ARHGAP35"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN056JYE|/analyses/ENCAN056JYE/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF991AML|/files/ENCFF991AML/}, {ENCFF010MMQ|/files/ENCFF010MMQ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.08 and a self consistency ratio of 1.19. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF205KOI|/files/ENCFF205KOI/}, {ENCFF967UPX|/files/ENCFF967UPX/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.08 and a self consistency ratio of 1.19. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR571BUF | float |
TF_ChIP-seq_ENCSR571BUF |
TF_ChIP-seq ENCSR571BUF [biosample_summary="Homo sapiens K562" and target="ARHGAP35"]
|
 |
[14.4, 1.47e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF010MMQ.bed.gz | 93.33 KB | c4d9d80170982a17ce764c0984b48c45 |
| ENCFF010MMQ.bed.gz.dvc | 99.0 B | 41c4bca24b9c31105f35abe8bbe8530e |
| ENCFF010MMQ.tabix.bed.gz | 60.74 KB | 6ad8dfc59ec5228298d615bacf6b1967 |
| ENCFF010MMQ.tabix.bed.gz.dvc | 105.0 B | e86c3fbbcd10ef888f142d817b0a7e30 |
| ENCFF010MMQ.tabix.bed.gz.tbi | 44.31 KB | 08ea61e31295fedc3b33d5752f3c1f7c |
| ENCFF010MMQ.tabix.bed.gz.tbi.dvc | 109.0 B | 759ab602f1956fd1b6d9128e922747d3 |
| genomic_resource.yaml | 3.03 KB | d28d5cdb7334e745f481c316e2508b0b |
| genomic_resource_original.yaml | 2.93 KB | b336d88388ec710e7551cea16ec52e21 |
| statistics/ |