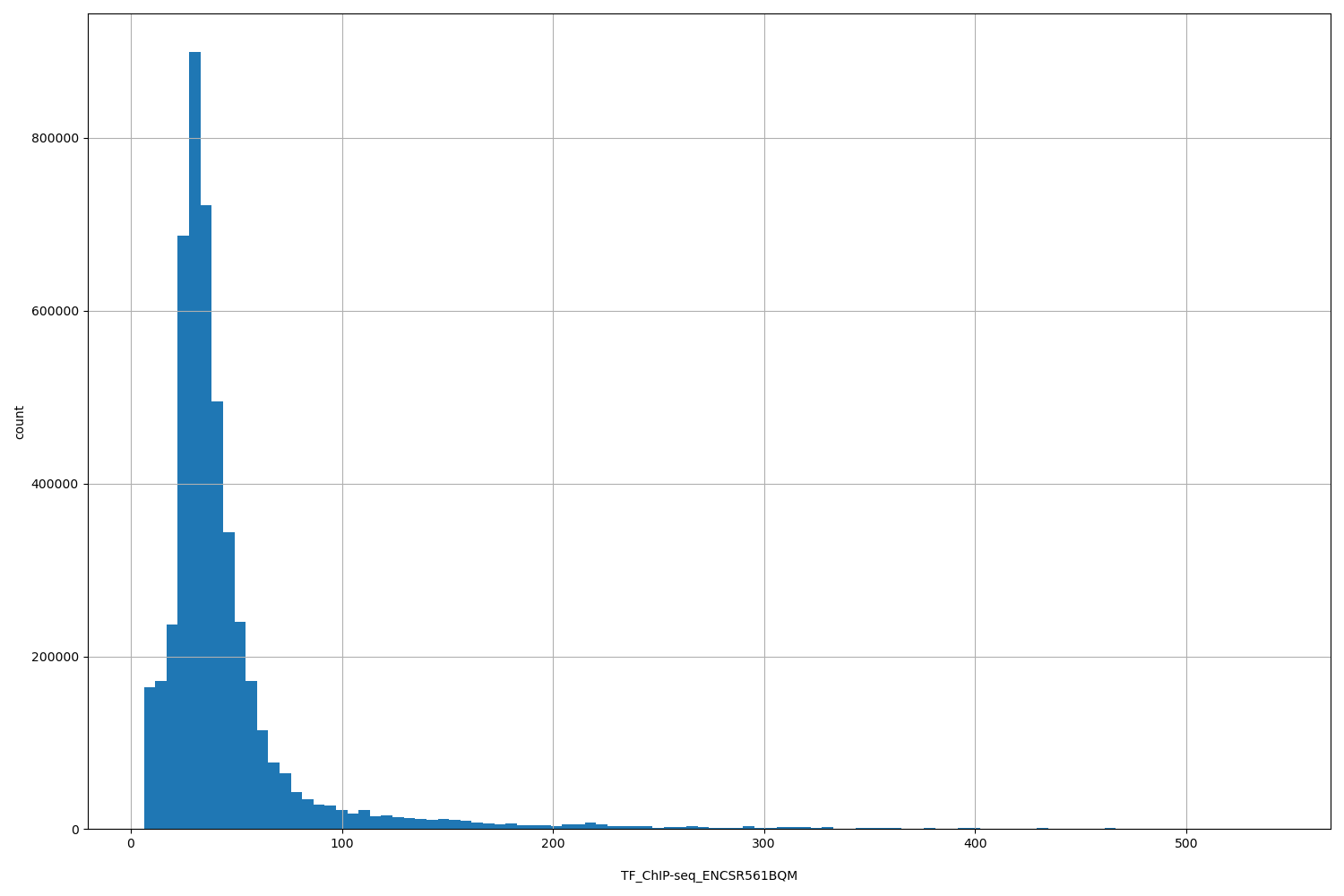
TF_ChIP-seq_ENCSR561BQM
| Id: | TF_ChIP-seq/ENCSR561BQM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR561BQM [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens SIX1" and target="SIX1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens SIX1 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN733ADU|/analyses/ENCAN733ADU/} has in progress subobject document {a462b83c-65ca-4c12-bfc5-e25c5db13831|/documents/a462b83c-65ca-4c12-bfc5-e25c5db13831/} audit_internal_action: Released analysis {ENCAN733ADU|/analyses/ENCAN733ADU/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF069QZX|/files/ENCFF069QZX/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.45 and a self consistency ratio of 3.97. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF587VYG|/files/ENCFF587VYG/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.45 and a self consistency ratio of 3.97. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR561BQM | float |
TF_ChIP-seq_ENCSR561BQM |
TF_ChIP-seq ENCSR561BQM [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens SIX1" and target="SIX1"]
|
 |
[6.2, 542] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF069QZX.bed.gz | 241.87 KB | bacf112145f7290b41aab0c683a36186 |
| ENCFF069QZX.bed.gz.dvc | 100.0 B | 7e6844b377e4255b34f91e798e2e8787 |
| ENCFF069QZX.tabix.bed.gz | 182.65 KB | b193663ab371e266d1ab8a14cb8edc2a |
| ENCFF069QZX.tabix.bed.gz.dvc | 106.0 B | f0e3d6892b62dc72af6aade7be573e07 |
| ENCFF069QZX.tabix.bed.gz.tbi | 99.88 KB | 9eee92f9cb06408d8a48fe8b16ec33f2 |
| ENCFF069QZX.tabix.bed.gz.tbi.dvc | 110.0 B | b4e693812269f9c103d2da87abf68520 |
| genomic_resource.yaml | 2.94 KB | 165a7bbab7ee43ebfd5698014be172e6 |
| genomic_resource_original.yaml | 2.78 KB | 90cb50ac376e75d1857d83e467ea6ddf |
| statistics/ |