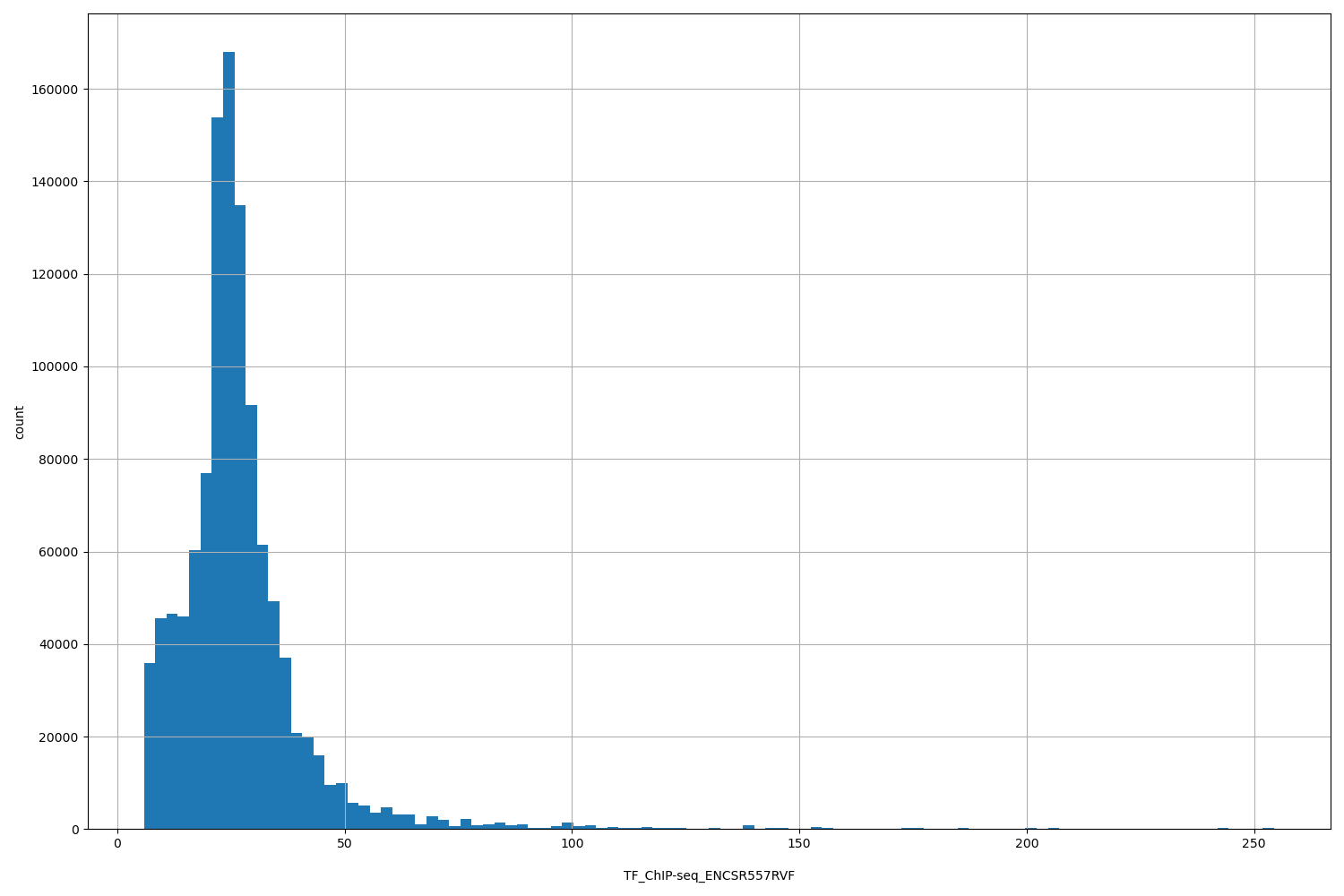
TF_ChIP-seq_ENCSR557RVF
| Id: | TF_ChIP-seq/ENCSR557RVF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR557RVF [biosamplesummary="Homo sapiens K562" and target="ZHX1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 3 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN279EUV|/analyses/ENCAN279EUV/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF899GZH|/files/ENCFF899GZH/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19589559 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZHX1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR557RVF | float |
TF_ChIP-seq_ENCSR557RVF |
TF_ChIP-seq ENCSR557RVF [biosample_summary="Homo sapiens K562" and target="ZHX1"]
|
 |
[5.83, 254] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF195ODE.bed.gz | 64.46 KB | a1e2f7cfe616efa0fd80d7f1210ba11f |
| ENCFF195ODE.bed.gz.dvc | 99.0 B | bd5686ec165a507d3a2581d0fa2a3867 |
| ENCFF195ODE.tabix.bed.gz | 45.3 KB | 766aaa4250a084969a45de614792cb54 |
| ENCFF195ODE.tabix.bed.gz.dvc | 105.0 B | 9279988a3c013c674eb42be243be2833 |
| ENCFF195ODE.tabix.bed.gz.tbi | 32.46 KB | 7c0d0920f0ed60936542b1ef15d940ae |
| ENCFF195ODE.tabix.bed.gz.tbi.dvc | 109.0 B | 75c8647fe44d5347238e501008d823d4 |
| genomic_resource.yaml | 1.83 KB | 0b3990961dbf9cb3de08dd34b7f2ed79 |
| genomic_resource_original.yaml | 1.74 KB | 539b907cd259b85e93698c66b1bbefd3 |
| statistics/ |