TF_ChIP-seq_ENCSR549NPZ
| Id: | TF_ChIP-seq/ENCSR549NPZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR549NPZ [biosamplesummary="Homo sapiens GM12878" and target="CBX3"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN327IDD|/analyses/ENCAN327IDD/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF177HNF|/files/ENCFF177HNF/} processed by ChIP-seq ENCODE3 hg19 pipeline has 18991913 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CBX3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
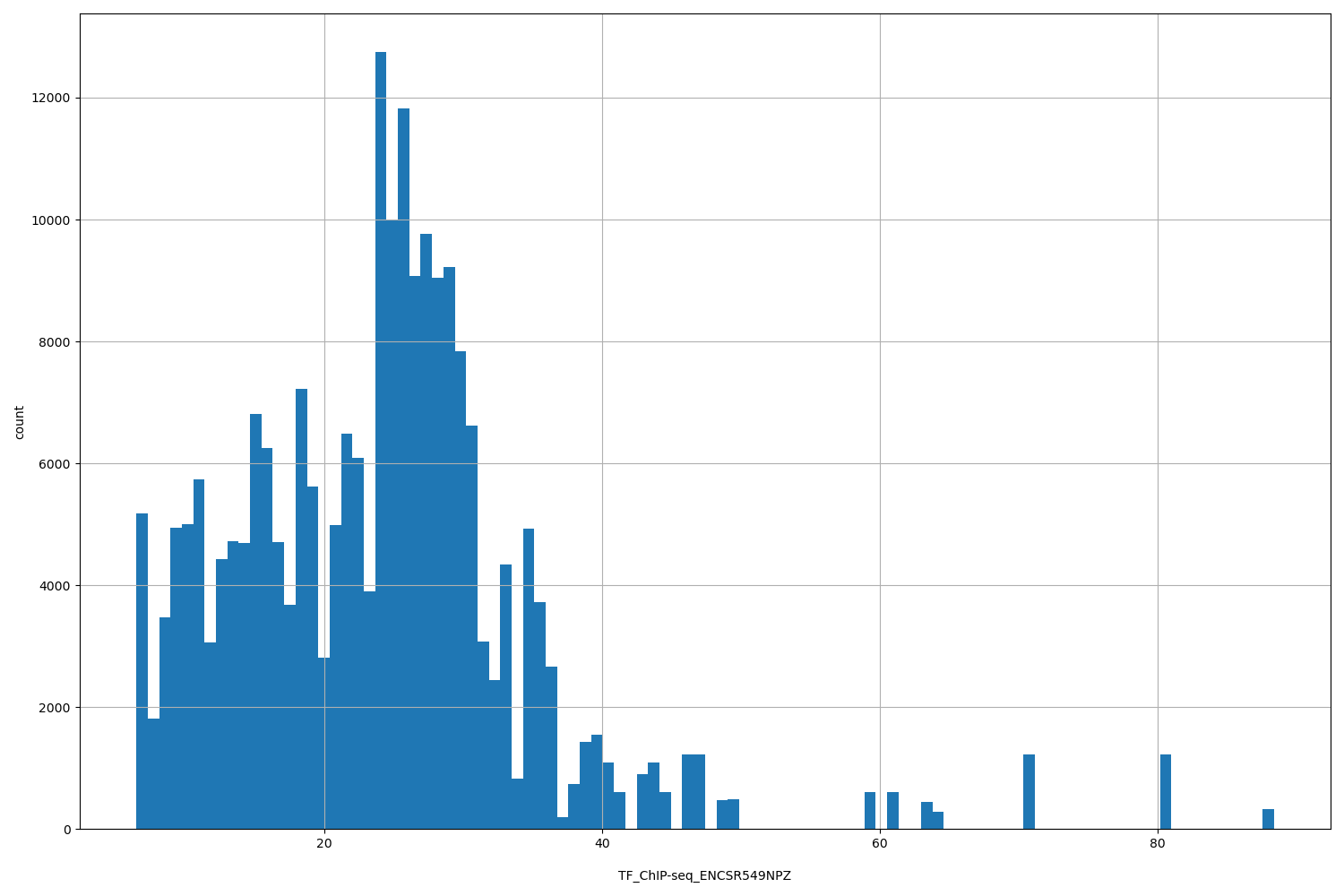
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR549NPZ | float |
TF_ChIP-seq_ENCSR549NPZ |
TF_ChIP-seq ENCSR549NPZ [biosample_summary="Homo sapiens GM12878" and target="CBX3"]
|
 |
[6.45, 88.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF141LEI.bed.gz | 8.32 KB | 58281cd2d3fdcbce6474dcc7abc61e32 |
| ENCFF141LEI.bed.gz.dvc | 98.0 B | a6191e0ed48ede1d996a3a2fdae4f50c |
| ENCFF141LEI.tabix.bed.gz | 6.42 KB | aaadb1375b1ccaaa14c660122a584671 |
| ENCFF141LEI.tabix.bed.gz.dvc | 104.0 B | 4b62280d6807809e504a69bebb176a53 |
| ENCFF141LEI.tabix.bed.gz.tbi | 7.42 KB | 08bc61a4115eff102bf91f16afa3c1ab |
| ENCFF141LEI.tabix.bed.gz.tbi.dvc | 108.0 B | db7778ca0609dcabff89d89eae109c7f |
| genomic_resource.yaml | 1.84 KB | f8f6b397aaec8c9024517755298f004c |
| genomic_resource_original.yaml | 1.74 KB | 177607391f05183db40d803d1a333a3e |
| statistics/ |