TF_ChIP-seq_ENCSR546KCN
| Id: | TF_ChIP-seq/ENCSR546KCN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR546KCN [biosamplesummary="Homo sapiens MCF-7 stably expressing FOSL2" and target="FOSL2"] |
| Description: |
status: archived biological_replicates: Rep 2, Rep 3 summary: stably expressing C-terminal eGFP-tagged FOSL2 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN400EJT|/analyses/ENCAN400EJT/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF121PXE|/files/ENCFF121PXE/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18936756 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting FOSL2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
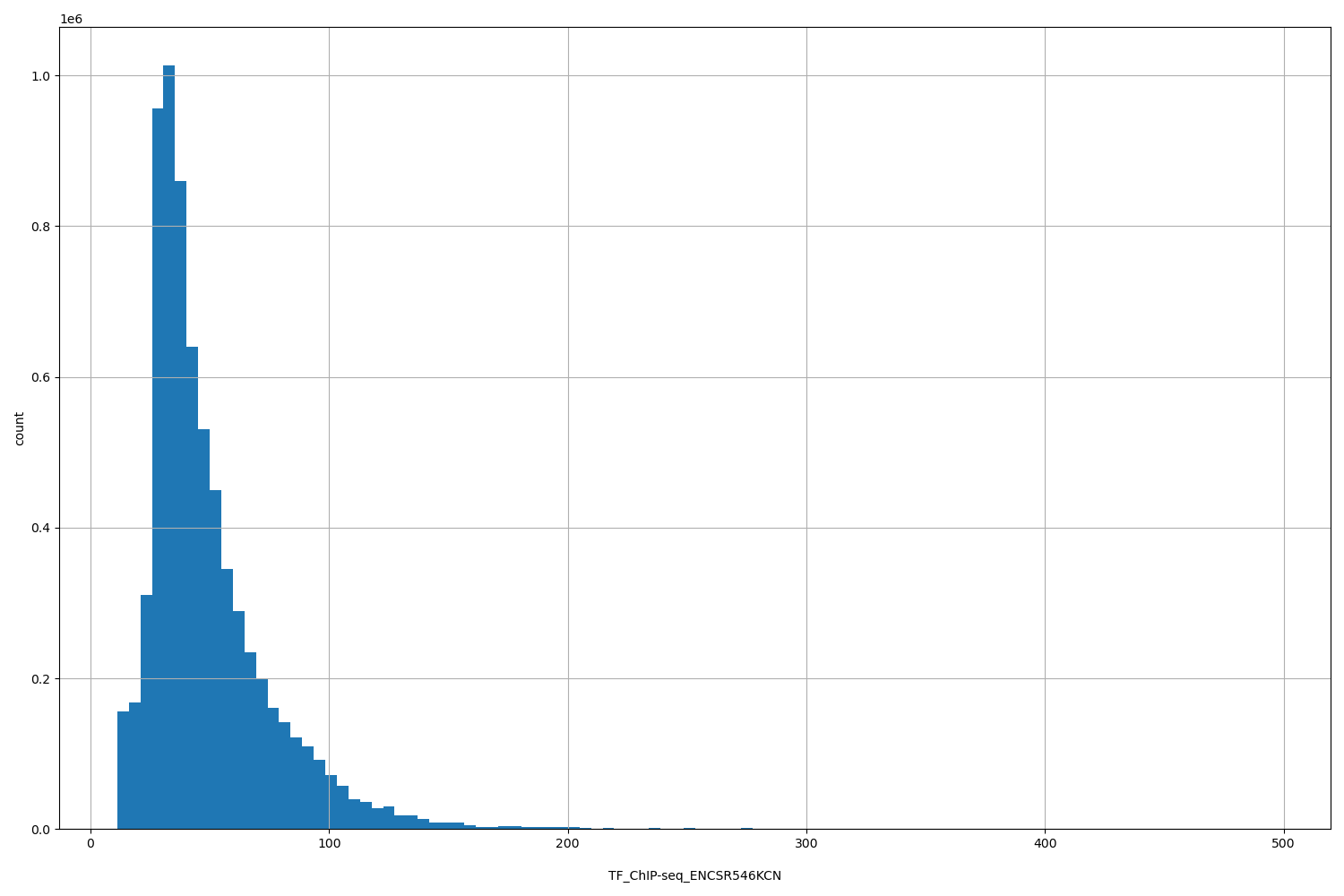
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR546KCN | float |
TF_ChIP-seq_ENCSR546KCN |
TF_ChIP-seq ENCSR546KCN [biosample_summary="Homo sapiens MCF-7 stably expressing FOSL2" and target="FOSL2"]
|
 |
[11.2, 495] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF998CZO.bed.gz | 248.7 KB | 94b565ffb2fb26b0290ddeebb4a495eb |
| ENCFF998CZO.bed.gz.dvc | 100.0 B | 8def2de67ae2d7d1fefeadfbf76fda64 |
| ENCFF998CZO.tabix.bed.gz | 190.93 KB | b70522666edbcea005007114041582ce |
| ENCFF998CZO.tabix.bed.gz.dvc | 106.0 B | bf17b00b6639aeba280ae75e9196de26 |
| ENCFF998CZO.tabix.bed.gz.tbi | 99.66 KB | 42423cac040e65e4de42217ebc495ac2 |
| ENCFF998CZO.tabix.bed.gz.tbi.dvc | 110.0 B | f5e1f7e04cb83a925af09fd83ffa610b |
| genomic_resource.yaml | 1.97 KB | d3015714d21e7a504ebfbc25e349f347 |
| genomic_resource_original.yaml | 1.86 KB | b32cb9975004953f04b7d6e51f34bff5 |
| statistics/ |