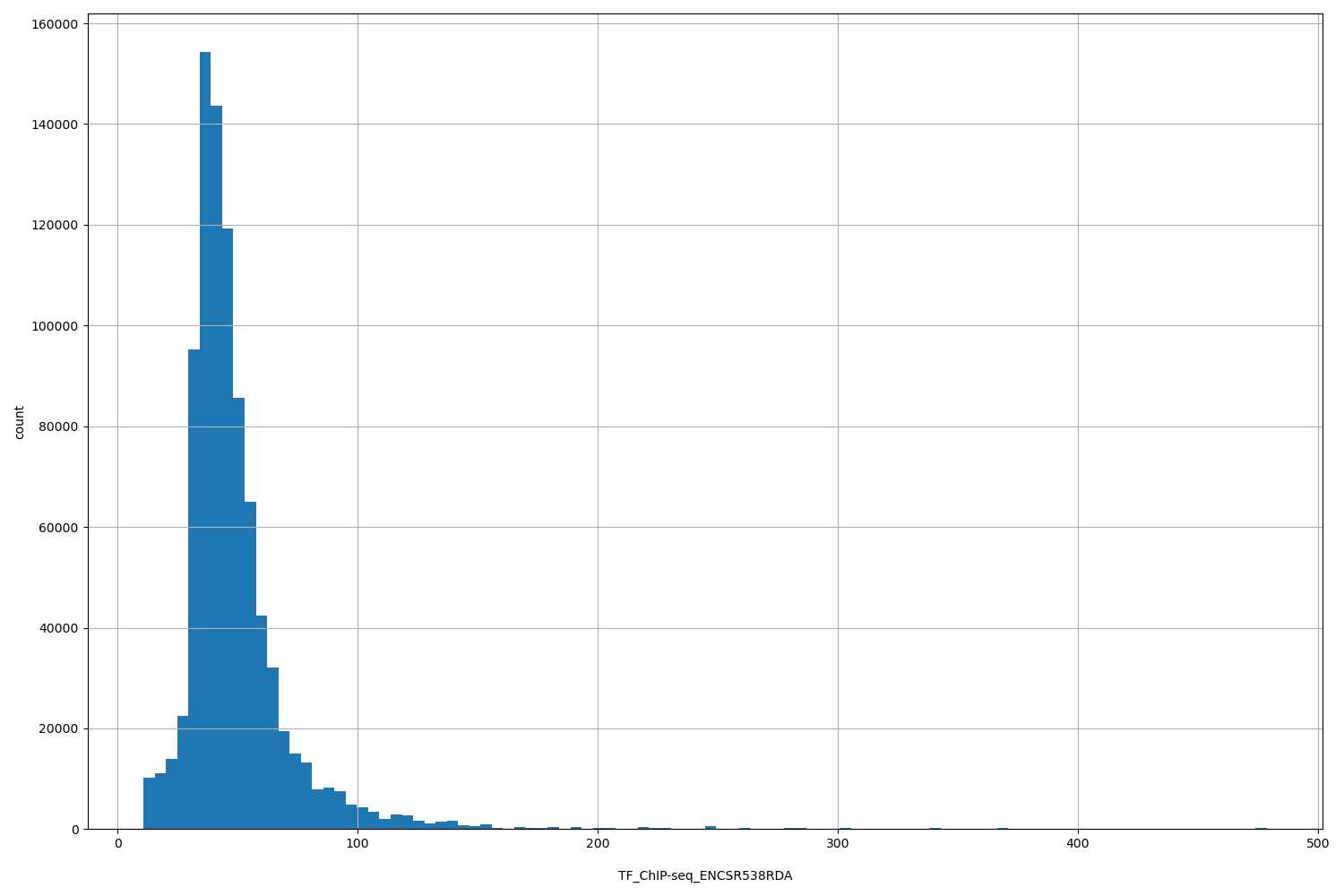
TF_ChIP-seq_ENCSR538RDA
| Id: | TF_ChIP-seq/ENCSR538RDA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR538RDA [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF560" and target="ZNF560"] |
| Description: |
status: released biological_replicates: Rep 2, Rep 3 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF560 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN387VWP|/analyses/ENCAN387VWP/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN387VWP|/analyses/ENCAN387VWP/} has in progress subobject document {c319166e-746d-4d6a-983f-8349593e7f3d|/documents/c319166e-746d-4d6a-983f-8349593e7f3d/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF173MMC|/files/ENCFF173MMC/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.20 and a self consistency ratio of 2.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF901CEW|/files/ENCFF901CEW/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.20 and a self consistency ratio of 2.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR538RDA | float |
TF_ChIP-seq_ENCSR538RDA |
TF_ChIP-seq ENCSR538RDA [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF560" and target="ZNF560"]
|
 |
[10.8, 479] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF173MMC.bed.gz | 66.16 KB | be54148cca63b8b9b7c75bd05a192e23 |
| ENCFF173MMC.bed.gz.dvc | 99.0 B | 03b95d8809ec2cf5d6997365cbf15a4a |
| ENCFF173MMC.tabix.bed.gz | 42.26 KB | 107ab69716879f4150896b6d2591452e |
| ENCFF173MMC.tabix.bed.gz.dvc | 105.0 B | f978f3622d029f695f997883cb4f1159 |
| ENCFF173MMC.tabix.bed.gz.tbi | 37.55 KB | cd3f70bea9bb99a2351c328ce2206e58 |
| ENCFF173MMC.tabix.bed.gz.tbi.dvc | 109.0 B | debb6a23aa9430e1e169e84b0b6b59e6 |
| genomic_resource.yaml | 3.07 KB | 0b33f87b31b5d10ebdca3cf6bb890eec |
| genomic_resource_original.yaml | 2.88 KB | 3051f6d83d624bd1eb7ced39594f5842 |
| statistics/ |