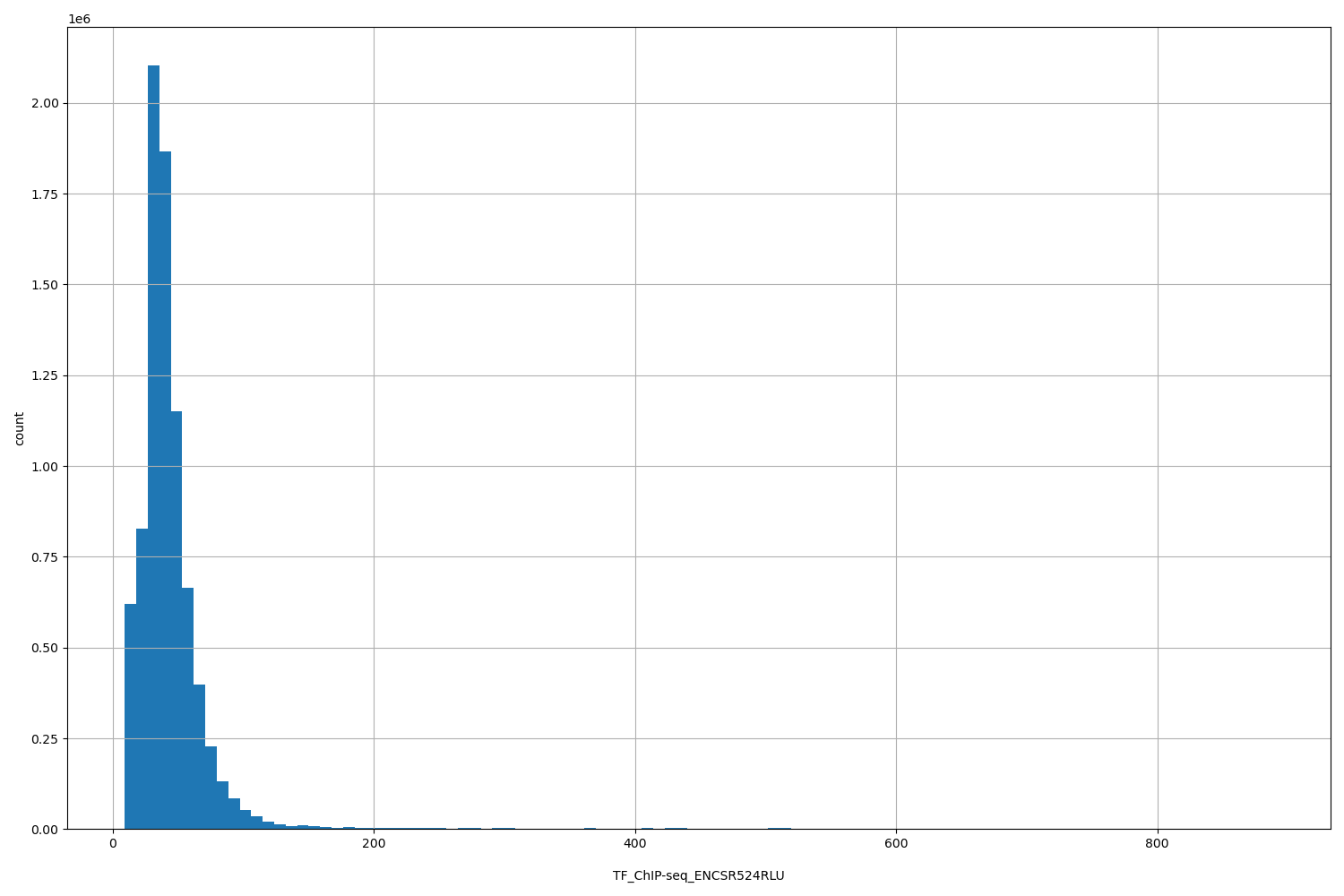
TF_ChIP-seq_ENCSR524RLU
| Id: | TF_ChIP-seq/ENCSR524RLU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR524RLU [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens RFXAP" and target="RFXAP"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens RFXAP output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN020FIE|/analyses/ENCAN020FIE/} has in progress subobject document {1b39924c-836c-4074-a332-8d8a00d15cc0|/documents/1b39924c-836c-4074-a332-8d8a00d15cc0/} audit_internal_action: Released analysis {ENCAN020FIE|/analyses/ENCAN020FIE/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF322CXC|/files/ENCFF322CXC/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.57 and a self consistency ratio of 2.39. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF359QOX|/files/ENCFF359QOX/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.57 and a self consistency ratio of 2.39. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR524RLU | float |
TF_ChIP-seq_ENCSR524RLU |
TF_ChIP-seq ENCSR524RLU [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens RFXAP" and target="RFXAP"]
|
 |
[9.4, 889] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF322CXC.bed.gz | 338.62 KB | 6c67c14c83c9ad380e561b803f5dbb5b |
| ENCFF322CXC.bed.gz.dvc | 100.0 B | 7973aa5b693923b9279612a921946d28 |
| ENCFF322CXC.tabix.bed.gz | 259.72 KB | c4de32233aaab534094c25106ecc21ee |
| ENCFF322CXC.tabix.bed.gz.dvc | 106.0 B | b6a6ceb9f273298c0e85e4762b4d6442 |
| ENCFF322CXC.tabix.bed.gz.tbi | 108.05 KB | a27b4cbd32d7fffd0dd8fce338bf8369 |
| ENCFF322CXC.tabix.bed.gz.tbi.dvc | 110.0 B | 14582bf97c3c91933c7801927c753249 |
| genomic_resource.yaml | 2.95 KB | e61cacfcae81883c7d1c94c49c12e9fa |
| genomic_resource_original.yaml | 2.79 KB | 0ffaa79f8f8c37ed7374cb952d0b2880 |
| statistics/ |